GPAW + WannierBerri: spinless bandstructure (diamond)
This tutorial demonstrates how to wannierize a spinless bandstructure of diamond
Prerequisites
Make sure you have installed the following packages:
pip install wannierberri gpaw ase irrep
Step 1: Gpaw calculation
self-consistent calculation
[2]:
import numpy as np
from ase import Atoms
from gpaw import GPAW, PW, MixerSum
from irrep.spacegroup import SpaceGroup
seed = "diamond"
[ ]:
a = 3.227
lattice = a * (np.ones((3, 3)) - np.eye(3)) / 2
positions = np.array([[0, 0, 0], [1, 1, 1]])/4
typeat = [1, 1]
atoms = Atoms(
"C2", cell=lattice, pbc=[1, 1, 1], scaled_positions=positions
)
calc = GPAW(
mode=PW(500),
xc="PBE",
symmetry={'symmorphic': False},
kpts={"size": [8, 8, 8], "gamma": True},
convergence={"density": 1e-6},
mixer=MixerSum(0.25, 8, 100),
txt=f"{seed}-scf.txt"
)
atoms.calc = calc
atoms.get_potential_energy()
calc.write(f"{seed}-scf.gpw", mode="all")
Non-self-consistent calculation in the irreducible Brillouin zone
Now we get the list of irreducible k-points (using irrep.SpaceGroup object) and perform a non-self-consistent calculation only on these k-points. We use irrep (instead of internal GPAW symmetrization), because it cann account for non-symmorphic symmetries, and also magnetic symmetries (see other tutorials)
[2]:
calc_scf = GPAW(f'{seed}-scf.gpw', txt=None)
sg = SpaceGroup.from_gpaw(calc_scf)
sg.show()
irred_kpt = sg.get_irreducible_kpoints_grid((4, 4, 4))
calc_nscf_irred = calc_scf.fixed_density(
kpts=irred_kpt,
symmetry={'symmorphic': False},
nbands=20,
convergence={'bands': 20},
txt=f'{seed}-nscf-irred.txt')
calc_nscf_irred.write(f'{seed}-nscf.gpw', mode='all')
typat used for spacegroup detection (accounting magmoms): [6 6]
---------- CRYSTAL STRUCTURE ----------
Cell vectors in angstroms:
Vectors of DFT cell
a0 = 0.0000 1.6135 1.6135
a1 = 1.6135 0.0000 1.6135
a2 = 1.6135 1.6135 0.0000
---------- SPACE GROUP -----------
Space group: Fd-3m1' (# 227.129)
Number of symmetries: 96 (mod. lattice translations)
### 1
rotation : | 1 0 0 |
| 0 1 0 |
| 0 0 1 |
gk = [kx, ky, kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.091821 -0.314892 0.944676] ; angle = 0 , inversion: False, time reversal: False
### 2
rotation : | 1 0 0 |
| 0 1 0 |
| 0 0 1 |
gk = [-kx, -ky, -kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.091821 -0.314892 0.944676] ; angle = 0 , inversion: False, time reversal: True
### 3
rotation : | 1 1 1 |
| 0 0 -1 |
| -1 0 0 |
gk = [ky, ky-kz, -kx+ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0. 0. 1.] ; angle = 1/2 pi, inversion: False, time reversal: False
### 4
rotation : | 1 1 1 |
| 0 0 -1 |
| -1 0 0 |
gk = [-ky, -ky+kz, kx-ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0. 0. 1.] ; angle = 1/2 pi, inversion: False, time reversal: True
### 5
rotation : | 0 1 0 |
| 1 0 0 |
| -1 -1 -1 |
gk = [ky-kz, kx-kz, -kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0. 0. -1.] ; angle = 1 pi, inversion: False, time reversal: False
### 6
rotation : | 0 1 0 |
| 1 0 0 |
| -1 -1 -1 |
gk = [-ky+kz, -kx+kz, kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0. 0. -1.] ; angle = 1 pi, inversion: False, time reversal: True
### 7
rotation : | 0 0 -1 |
| 1 1 1 |
| 0 -1 0 |
gk = [kx-kz, kx, kx-ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0. 0. 1.] ; angle = -1/2 pi, inversion: False, time reversal: False
### 8
rotation : | 0 0 -1 |
| 1 1 1 |
| 0 -1 0 |
gk = [-kx+kz, -kx, -kx+ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0. 0. 1.] ; angle = -1/2 pi, inversion: False, time reversal: True
### 9
rotation : | -1 -1 -1 |
| 0 0 1 |
| 0 1 0 |
gk = [-kx, -kx+kz, -kx+ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [1. 0. 0.] ; angle = 1 pi, inversion: False, time reversal: False
### 10
rotation : | -1 -1 -1 |
| 0 0 1 |
| 0 1 0 |
gk = [kx, kx-kz, kx-ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [1. 0. 0.] ; angle = 1 pi, inversion: False, time reversal: True
### 11
rotation : | 0 -1 0 |
| -1 0 0 |
| 0 0 -1 |
gk = [-ky, -kx, -kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.707107 -0.707107 -0. ] ; angle = 1 pi, inversion: False, time reversal: False
### 12
rotation : | 0 -1 0 |
| -1 0 0 |
| 0 0 -1 |
gk = [ky, kx, kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.707107 -0.707107 -0. ] ; angle = 1 pi, inversion: False, time reversal: True
### 13
rotation : | 0 0 1 |
| -1 -1 -1 |
| 1 0 0 |
gk = [-ky+kz, -ky, kx-ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0. 1. -0.] ; angle = 1 pi, inversion: False, time reversal: False
### 14
rotation : | 0 0 1 |
| -1 -1 -1 |
| 1 0 0 |
gk = [ky-kz, ky, -kx+ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0. 1. -0.] ; angle = 1 pi, inversion: False, time reversal: True
### 15
rotation : | -1 0 0 |
| 0 -1 0 |
| 1 1 1 |
gk = [-kx+kz, -ky+kz, kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.707107 0.707107 -0. ] ; angle = 1 pi, inversion: False, time reversal: False
### 16
rotation : | -1 0 0 |
| 0 -1 0 |
| 1 1 1 |
gk = [kx-kz, ky-kz, -kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.707107 0.707107 -0. ] ; angle = 1 pi, inversion: False, time reversal: True
### 17
rotation : | 0 0 1 |
| 1 0 0 |
| 0 1 0 |
gk = [kz, kx, ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [0.57735 0.57735 0.57735] ; angle = 2/3 pi, inversion: False, time reversal: False
### 18
rotation : | 0 0 1 |
| 1 0 0 |
| 0 1 0 |
gk = [-kz, -kx, -ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [0.57735 0.57735 0.57735] ; angle = 2/3 pi, inversion: False, time reversal: True
### 19
rotation : | -1 0 0 |
| 1 1 1 |
| 0 0 -1 |
gk = [-kx+ky, ky, ky-kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0.707107 0. 0.707107] ; angle = 1 pi, inversion: False, time reversal: False
### 20
rotation : | -1 0 0 |
| 1 1 1 |
| 0 0 -1 |
gk = [kx-ky, -ky, -ky+kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0.707107 0. 0.707107] ; angle = 1 pi, inversion: False, time reversal: True
### 21
rotation : | -1 -1 -1 |
| 0 1 0 |
| 1 0 0 |
gk = [-kz, ky-kz, kx-kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 0.57735 -0.57735] ; angle = 2/3 pi, inversion: False, time reversal: False
### 22
rotation : | -1 -1 -1 |
| 0 1 0 |
| 1 0 0 |
gk = [kz, -ky+kz, -kx+kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 0.57735 -0.57735] ; angle = 2/3 pi, inversion: False, time reversal: True
### 23
rotation : | 0 -1 0 |
| 0 0 -1 |
| 1 1 1 |
gk = [kx-ky, kx-kz, kx]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0. 1. 0.] ; angle = 1/2 pi, inversion: False, time reversal: False
### 24
rotation : | 0 -1 0 |
| 0 0 -1 |
| 1 1 1 |
gk = [-kx+ky, -kx+kz, -kx]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0. 1. 0.] ; angle = 1/2 pi, inversion: False, time reversal: True
### 25
rotation : | 0 1 0 |
| -1 -1 -1 |
| 0 0 1 |
gk = [-kx+ky, -kx, -kx+kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.57735 0.57735 -0.57735] ; angle = -2/3 pi, inversion: False, time reversal: False
### 26
rotation : | 0 1 0 |
| -1 -1 -1 |
| 0 0 1 |
gk = [kx-ky, kx, kx-kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.57735 0.57735 -0.57735] ; angle = -2/3 pi, inversion: False, time reversal: True
### 27
rotation : | 0 0 -1 |
| 0 -1 0 |
| -1 0 0 |
gk = [-kz, -ky, -kx]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-0.707107 0. 0.707107] ; angle = 1 pi, inversion: False, time reversal: False
### 28
rotation : | 0 0 -1 |
| 0 -1 0 |
| -1 0 0 |
gk = [kz, ky, kx]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-0.707107 0. 0.707107] ; angle = 1 pi, inversion: False, time reversal: True
### 29
rotation : | 1 0 0 |
| 0 0 1 |
| -1 -1 -1 |
gk = [kx-ky, -ky+kz, -ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 0.57735 0.57735] ; angle = -2/3 pi, inversion: False, time reversal: False
### 30
rotation : | 1 0 0 |
| 0 0 1 |
| -1 -1 -1 |
gk = [-kx+ky, ky-kz, ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 0.57735 0.57735] ; angle = -2/3 pi, inversion: False, time reversal: True
### 31
rotation : | 1 1 1 |
| -1 0 0 |
| 0 -1 0 |
gk = [kz, -kx+kz, -ky+kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0. -1. 0.] ; angle = 1/2 pi, inversion: False, time reversal: False
### 32
rotation : | 1 1 1 |
| -1 0 0 |
| 0 -1 0 |
gk = [-kz, kx-kz, ky-kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0. -1. 0.] ; angle = 1/2 pi, inversion: False, time reversal: True
### 33
rotation : | 0 1 0 |
| 0 0 1 |
| 1 0 0 |
gk = [ky, kz, kx]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 -0.57735 -0.57735] ; angle = 2/3 pi, inversion: False, time reversal: False
### 34
rotation : | 0 1 0 |
| 0 0 1 |
| 1 0 0 |
gk = [-ky, -kz, -kx]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 -0.57735 -0.57735] ; angle = 2/3 pi, inversion: False, time reversal: True
### 35
rotation : | 0 0 -1 |
| -1 0 0 |
| 1 1 1 |
gk = [ky-kz, -kx+ky, ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-1. -0. 0.] ; angle = 1/2 pi, inversion: False, time reversal: False
### 36
rotation : | 0 0 -1 |
| -1 0 0 |
| 1 1 1 |
gk = [-ky+kz, kx-ky, -ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-1. -0. 0.] ; angle = 1/2 pi, inversion: False, time reversal: True
### 37
rotation : | 1 0 0 |
| -1 -1 -1 |
| 0 1 0 |
gk = [kx-kz, -kz, ky-kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 0.57735 0.57735] ; angle = 2/3 pi, inversion: False, time reversal: False
### 38
rotation : | 1 0 0 |
| -1 -1 -1 |
| 0 1 0 |
gk = [-kx+kz, kz, -ky+kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 0.57735 0.57735] ; angle = 2/3 pi, inversion: False, time reversal: True
### 39
rotation : | 1 1 1 |
| 0 -1 0 |
| 0 0 -1 |
gk = [kx, kx-ky, kx-kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0. -0.707107 -0.707107] ; angle = 1 pi, inversion: False, time reversal: False
### 40
rotation : | 1 1 1 |
| 0 -1 0 |
| 0 0 -1 |
gk = [-kx, -kx+ky, -kx+kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0. -0.707107 -0.707107] ; angle = 1 pi, inversion: False, time reversal: True
### 41
rotation : | 0 0 1 |
| 0 1 0 |
| -1 -1 -1 |
gk = [-kx+kz, -kx+ky, -kx]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.57735 -0.57735 0.57735] ; angle = 2/3 pi, inversion: False, time reversal: False
### 42
rotation : | 0 0 1 |
| 0 1 0 |
| -1 -1 -1 |
gk = [kx-kz, kx-ky, kx]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.57735 -0.57735 0.57735] ; angle = 2/3 pi, inversion: False, time reversal: True
### 43
rotation : | -1 0 0 |
| 0 0 -1 |
| 0 -1 0 |
gk = [-kx, -kz, -ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0. -0.707107 0.707107] ; angle = 1 pi, inversion: False, time reversal: False
### 44
rotation : | -1 0 0 |
| 0 0 -1 |
| 0 -1 0 |
gk = [kx, kz, ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0. -0.707107 0.707107] ; angle = 1 pi, inversion: False, time reversal: True
### 45
rotation : | -1 -1 -1 |
| 1 0 0 |
| 0 0 1 |
gk = [-ky, kx-ky, -ky+kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 -0.57735 0.57735] ; angle = -2/3 pi, inversion: False, time reversal: False
### 46
rotation : | -1 -1 -1 |
| 1 0 0 |
| 0 0 1 |
gk = [ky, -kx+ky, ky-kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.57735 -0.57735 0.57735] ; angle = -2/3 pi, inversion: False, time reversal: True
### 47
rotation : | 0 -1 0 |
| 1 1 1 |
| -1 0 0 |
gk = [-ky+kz, kz, -kx+kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-1. 0. 0.] ; angle = -1/2 pi, inversion: False, time reversal: False
### 48
rotation : | 0 -1 0 |
| 1 1 1 |
| -1 0 0 |
gk = [ky-kz, -kz, kx-kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-1. 0. 0.] ; angle = -1/2 pi, inversion: False, time reversal: True
### 49
rotation : | -1 0 0 |
| 0 -1 0 |
| 0 0 -1 |
gk = [-kx, -ky, -kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-0.091821 -0.314892 0.944676] ; angle = 0 , inversion: True, time reversal: False
### 50
rotation : | -1 0 0 |
| 0 -1 0 |
| 0 0 -1 |
gk = [kx, ky, kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-0.091821 -0.314892 0.944676] ; angle = 0 , inversion: True, time reversal: True
### 51
rotation : | -1 -1 -1 |
| 0 0 1 |
| 1 0 0 |
gk = [-ky, -ky+kz, kx-ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [0. 0. 1.] ; angle = 1/2 pi, inversion: True, time reversal: False
### 52
rotation : | -1 -1 -1 |
| 0 0 1 |
| 1 0 0 |
gk = [ky, ky-kz, -kx+ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [0. 0. 1.] ; angle = 1/2 pi, inversion: True, time reversal: True
### 53
rotation : | 0 -1 0 |
| -1 0 0 |
| 1 1 1 |
gk = [-ky+kz, -kx+kz, kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0. -0. 1.] ; angle = 1 pi, inversion: True, time reversal: False
### 54
rotation : | 0 -1 0 |
| -1 0 0 |
| 1 1 1 |
gk = [ky-kz, kx-kz, -kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0. -0. 1.] ; angle = 1 pi, inversion: True, time reversal: True
### 55
rotation : | 0 0 1 |
| -1 -1 -1 |
| 0 1 0 |
gk = [-kx+kz, -kx, -kx+ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [0. 0. 1.] ; angle = -1/2 pi, inversion: True, time reversal: False
### 56
rotation : | 0 0 1 |
| -1 -1 -1 |
| 0 1 0 |
gk = [kx-kz, kx, kx-ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [0. 0. 1.] ; angle = -1/2 pi, inversion: True, time reversal: True
### 57
rotation : | 1 1 1 |
| 0 0 -1 |
| 0 -1 0 |
gk = [kx, kx-kz, kx-ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [1. 0. 0.] ; angle = 1 pi, inversion: True, time reversal: False
### 58
rotation : | 1 1 1 |
| 0 0 -1 |
| 0 -1 0 |
gk = [-kx, -kx+kz, -kx+ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [1. 0. 0.] ; angle = 1 pi, inversion: True, time reversal: True
### 59
rotation : | 0 1 0 |
| 1 0 0 |
| 0 0 1 |
gk = [ky, kx, kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.707107 -0.707107 -0. ] ; angle = 1 pi, inversion: True, time reversal: False
### 60
rotation : | 0 1 0 |
| 1 0 0 |
| 0 0 1 |
gk = [-ky, -kx, -kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.707107 -0.707107 -0. ] ; angle = 1 pi, inversion: True, time reversal: True
### 61
rotation : | 0 0 -1 |
| 1 1 1 |
| -1 0 0 |
gk = [ky-kz, ky, -kx+ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0. 1. 0.] ; angle = 1 pi, inversion: True, time reversal: False
### 62
rotation : | 0 0 -1 |
| 1 1 1 |
| -1 0 0 |
gk = [-ky+kz, -ky, kx-ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0. 1. 0.] ; angle = 1 pi, inversion: True, time reversal: True
### 63
rotation : | 1 0 0 |
| 0 1 0 |
| -1 -1 -1 |
gk = [kx-kz, ky-kz, -kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.707107 -0.707107 0. ] ; angle = 1 pi, inversion: True, time reversal: False
### 64
rotation : | 1 0 0 |
| 0 1 0 |
| -1 -1 -1 |
gk = [-kx+kz, -ky+kz, kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.707107 -0.707107 0. ] ; angle = 1 pi, inversion: True, time reversal: True
### 65
rotation : | 0 0 -1 |
| -1 0 0 |
| 0 -1 0 |
gk = [-kz, -kx, -ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0.57735 0.57735 0.57735] ; angle = 2/3 pi, inversion: True, time reversal: False
### 66
rotation : | 0 0 -1 |
| -1 0 0 |
| 0 -1 0 |
gk = [kz, kx, ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0.57735 0.57735 0.57735] ; angle = 2/3 pi, inversion: True, time reversal: True
### 67
rotation : | 1 0 0 |
| -1 -1 -1 |
| 0 0 1 |
gk = [kx-ky, -ky, -ky+kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.707107 0. -0.707107] ; angle = 1 pi, inversion: True, time reversal: False
### 68
rotation : | 1 0 0 |
| -1 -1 -1 |
| 0 0 1 |
gk = [-kx+ky, ky, ky-kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-0.707107 0. -0.707107] ; angle = 1 pi, inversion: True, time reversal: True
### 69
rotation : | 1 1 1 |
| 0 -1 0 |
| -1 0 0 |
gk = [kz, -ky+kz, -kx+kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 -0.57735 0.57735] ; angle = -2/3 pi, inversion: True, time reversal: False
### 70
rotation : | 1 1 1 |
| 0 -1 0 |
| -1 0 0 |
gk = [-kz, ky-kz, kx-kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 -0.57735 0.57735] ; angle = -2/3 pi, inversion: True, time reversal: True
### 71
rotation : | 0 1 0 |
| 0 0 1 |
| -1 -1 -1 |
gk = [-kx+ky, -kx+kz, -kx]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [0. 1. 0.] ; angle = 1/2 pi, inversion: True, time reversal: False
### 72
rotation : | 0 1 0 |
| 0 0 1 |
| -1 -1 -1 |
gk = [kx-ky, kx-kz, kx]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [0. 1. 0.] ; angle = 1/2 pi, inversion: True, time reversal: True
### 73
rotation : | 0 -1 0 |
| 1 1 1 |
| 0 0 -1 |
gk = [kx-ky, kx, kx-kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 0.57735 -0.57735] ; angle = -2/3 pi, inversion: True, time reversal: False
### 74
rotation : | 0 -1 0 |
| 1 1 1 |
| 0 0 -1 |
gk = [-kx+ky, -kx, -kx+kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 0.57735 -0.57735] ; angle = -2/3 pi, inversion: True, time reversal: True
### 75
rotation : | 0 0 1 |
| 0 1 0 |
| 1 0 0 |
gk = [kz, ky, kx]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.707107 0. -0.707107] ; angle = 1 pi, inversion: True, time reversal: False
### 76
rotation : | 0 0 1 |
| 0 1 0 |
| 1 0 0 |
gk = [-kz, -ky, -kx]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0.707107 0. -0.707107] ; angle = 1 pi, inversion: True, time reversal: True
### 77
rotation : | -1 0 0 |
| 0 0 -1 |
| 1 1 1 |
gk = [-kx+ky, ky-kz, ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-0.57735 0.57735 0.57735] ; angle = -2/3 pi, inversion: True, time reversal: False
### 78
rotation : | -1 0 0 |
| 0 0 -1 |
| 1 1 1 |
gk = [kx-ky, -ky+kz, -ky]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [-0.57735 0.57735 0.57735] ; angle = -2/3 pi, inversion: True, time reversal: True
### 79
rotation : | -1 -1 -1 |
| 1 0 0 |
| 0 1 0 |
gk = [-kz, kx-kz, ky-kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0. -1. 0.] ; angle = 1/2 pi, inversion: True, time reversal: False
### 80
rotation : | -1 -1 -1 |
| 1 0 0 |
| 0 1 0 |
gk = [kz, -kx+kz, -ky+kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0. -1. 0.] ; angle = 1/2 pi, inversion: True, time reversal: True
### 81
rotation : | 0 -1 0 |
| 0 0 -1 |
| -1 0 0 |
gk = [-ky, -kz, -kx]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0.57735 0.57735 0.57735] ; angle = -2/3 pi, inversion: True, time reversal: False
### 82
rotation : | 0 -1 0 |
| 0 0 -1 |
| -1 0 0 |
gk = [ky, kz, kx]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [0.57735 0.57735 0.57735] ; angle = -2/3 pi, inversion: True, time reversal: True
### 83
rotation : | 0 0 1 |
| 1 0 0 |
| -1 -1 -1 |
gk = [-ky+kz, kx-ky, -ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-1. -0. 0.] ; angle = 1/2 pi, inversion: True, time reversal: False
### 84
rotation : | 0 0 1 |
| 1 0 0 |
| -1 -1 -1 |
gk = [ky-kz, -kx+ky, ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-1. -0. 0.] ; angle = 1/2 pi, inversion: True, time reversal: True
### 85
rotation : | -1 0 0 |
| 1 1 1 |
| 0 -1 0 |
gk = [-kx+kz, kz, -ky+kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 -0.57735 -0.57735] ; angle = -2/3 pi, inversion: True, time reversal: False
### 86
rotation : | -1 0 0 |
| 1 1 1 |
| 0 -1 0 |
gk = [kx-kz, -kz, ky-kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 -0.57735 -0.57735] ; angle = -2/3 pi, inversion: True, time reversal: True
### 87
rotation : | -1 -1 -1 |
| 0 1 0 |
| 0 0 1 |
gk = [-kx, -kx+ky, -kx+kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0. -0.707107 -0.707107] ; angle = 1 pi, inversion: True, time reversal: False
### 88
rotation : | -1 -1 -1 |
| 0 1 0 |
| 0 0 1 |
gk = [kx, kx-ky, kx-kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0. -0.707107 -0.707107] ; angle = 1 pi, inversion: True, time reversal: True
### 89
rotation : | 0 0 -1 |
| 0 -1 0 |
| 1 1 1 |
gk = [kx-kz, kx-ky, kx]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 -0.57735 0.57735] ; angle = 2/3 pi, inversion: True, time reversal: False
### 90
rotation : | 0 0 -1 |
| 0 -1 0 |
| 1 1 1 |
gk = [-kx+kz, -kx+ky, -kx]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 -0.57735 0.57735] ; angle = 2/3 pi, inversion: True, time reversal: True
### 91
rotation : | 1 0 0 |
| 0 0 1 |
| 0 1 0 |
gk = [kx, kz, ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0. -0.707107 0.707107] ; angle = 1 pi, inversion: True, time reversal: False
### 92
rotation : | 1 0 0 |
| 0 0 1 |
| 0 1 0 |
gk = [-kx, -kz, -ky]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [ 0. -0.707107 0.707107] ; angle = 1 pi, inversion: True, time reversal: True
### 93
rotation : | 1 1 1 |
| -1 0 0 |
| 0 0 -1 |
gk = [ky, -kx+ky, ky-kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 0.57735 -0.57735] ; angle = 2/3 pi, inversion: True, time reversal: False
### 94
rotation : | 1 1 1 |
| -1 0 0 |
| 0 0 -1 |
gk = [-ky, kx-ky, -ky+kz]
translation : [ 0.2500 0.2500 0.2500 ]
axis: [ 0.57735 0.57735 -0.57735] ; angle = 2/3 pi, inversion: True, time reversal: True
### 95
rotation : | 0 1 0 |
| -1 -1 -1 |
| 1 0 0 |
gk = [ky-kz, -kz, kx-kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-1. 0. 0.] ; angle = -1/2 pi, inversion: True, time reversal: False
### 96
rotation : | 0 1 0 |
| -1 -1 -1 |
| 1 0 0 |
gk = [-ky+kz, kz, -kx+kz]
translation : [ 0.0000 0.0000 0.0000 ]
axis: [-1. 0. 0.] ; angle = -1/2 pi, inversion: True, time reversal: True
Compute the dft bandstructure along a high-symmetry path
This is done to compare with the wannierized bandstructure later.
[3]:
calc_bands = GPAW(f"{seed}-scf.gpw",).fixed_density(
nbands=16,
symmetry='off',
kpts={'path': 'WLGXWKG', 'npoints': 60},
convergence={'bands': 8})
calc_bands.write(f"{seed}-bands.gpw", mode="all")
___ ___ ___ _ _ _
| | |_ | | | |
| | | | | . | | | |
|__ | _|___|_____| 25.7.0
|___|_|
User: stepan@C3MP-C019202L
Date: Thu Apr 2 12:57:27 2026
Arch: x86_64
Pid: 15866
CWD: /home/stepan/github/WannierBerri-tutorial/tutorials/8_GPAW/1.diamond
Python: 3.12.12
gpaw: /home/stepan/github/WannierBerri-tutorial/.conda/lib/python3.12/site-packages/gpaw
_gpaw: /home/stepan/github/WannierBerri-tutorial/.conda/lib/python3.12/site-packages/
_gpaw.cpython-312-x86_64-linux-gnu.so
ase: /home/stepan/github/WannierBerri-tutorial/.conda/lib/python3.12/site-packages/ase (version 3.26.0)
numpy: /home/stepan/github/WannierBerri-tutorial/.conda/lib/python3.12/site-packages/numpy (version 2.3.4)
scipy: /home/stepan/github/WannierBerri-tutorial/.conda/lib/python3.12/site-packages/scipy (version 1.16.2)
libxc: 5.2.3
units: Angstrom and eV
cores: 1
OpenMP: False
OMP_NUM_THREADS: 1
Input parameters:
convergence: {bands: 8}
gpts: [12 12 12]
kpts: {npoints: 60,
path: WLGXWKG}
mixer: {backend: pulay,
beta: 0.25,
method: sum,
nmaxold: 8,
weight: 100}
mode: {ecut: 500.0,
name: pw}
nbands: 16
symmetry: off
xc: PBE
Initialize ...
species:
C:
name: Carbon
id: 4aa54d4b901d75f77cc0ea3eec22967b
Z: 6.0
valence: 4
core: 2
charge: 0.0
file: /home/stepan/gpaw-data/gpaw-setups-24.11.0/C.PBE.gz
compensation charges: {type: gauss,
rc: 0.20,
lmax: 2}
cutoffs: {filter: 1.14,
core: 1.14}
projectors:
# energy rcut
- 2s(2.00) -13.751 0.635
- 2p(2.00) -5.284 0.635
- s 13.461 0.635
- p 21.927 0.635
- d 0.000 0.635
# Using partial waves for C as LCAO basis
Reference energy: -2055.256231 # eV
Spin-paired calculation
Convergence criteria:
Maximum [total energy] change in last 3 cyles: 0.0005 eV / valence electron
Maximum integral of absolute [dens]ity change: 0.0001 electrons / valence electron
Maximum integral of absolute [eigenst]ate change: 4e-08 eV^2 / valence electron
Maximum number of scf [iter]ations: 333
(Square brackets indicate name in SCF output, whereas a 'c' in
the SCF output indicates the quantity has converged.)
Symmetries present (total): 1
( 1 0 0)
( 0 1 0)
( 0 0 1)
60 k-points
60 k-points in the irreducible part of the Brillouin zone
k-points in crystal coordinates weights
0: 0.50000000 0.25000000 0.75000000 0.01666667
1: 0.50000000 0.28125000 0.71875000 0.01666667
2: 0.50000000 0.31250000 0.68750000 0.01666667
3: 0.50000000 0.34375000 0.65625000 0.01666667
4: 0.50000000 0.37500000 0.62500000 0.01666667
5: 0.50000000 0.40625000 0.59375000 0.01666667
6: 0.50000000 0.43750000 0.56250000 0.01666667
7: 0.50000000 0.46875000 0.53125000 0.01666667
8: 0.50000000 0.50000000 0.50000000 0.01666667
9: 0.45454545 0.45454545 0.45454545 0.01666667
...
59: 0.00000000 0.00000000 0.00000000 0.01666667
Wave functions: Plane wave expansion
Cutoff energy: 500.000 eV
Number of coefficients (min, max): 181, 222
Pulay-stress correction: 0.000000 eV/Ang^3 (de/decut=0.000000)
Using Numpy's FFT
ScaLapack parameters: grid=1x1, blocksize=None
Wavefunction extrapolation:
Improved wavefunction reuse through dual PAW basis
Occupation numbers: Fermi-Dirac:
width: 0.1000 # eV
Eigensolver
Davidson(niter=2)
Densities:
Coarse grid: 12*12*12 grid
Fine grid: 24*24*24 grid
Total Charge: 0.000000
Density mixing:
Method: separate
Backend: pulay
Linear mixing parameter: 0.25
old densities: 8
Damping of long wavelength oscillations: 100
Hamiltonian:
XC and Coulomb potentials evaluated on a 24*24*24 grid
Using the PBE Exchange-Correlation functional
External potential:
NoExternalPotential
XC parameters: PBE with 2 nearest neighbor stencil
Memory estimate:
Process memory now: 268.07 MiB
Calculator: 6.37 MiB
Density: 0.78 MiB
Arrays: 0.34 MiB
Localized functions: 0.23 MiB
Mixer: 0.21 MiB
Hamiltonian: 0.23 MiB
Arrays: 0.22 MiB
XC: 0.00 MiB
Poisson: 0.00 MiB
vbar: 0.00 MiB
Wavefunctions: 5.36 MiB
Arrays psit_nG: 3.25 MiB
Eigensolver: 0.11 MiB
Projections: 0.38 MiB
Projectors: 1.35 MiB
PW-descriptor: 0.27 MiB
Total number of cores used: 1
Number of atoms: 2
Number of atomic orbitals: 8
Number of bands in calculation: 16
Number of valence electrons: 8
Bands to converge: 8
... initialized
Initializing position-dependent things.
Creating initial wave functions:
8 bands from LCAO basis set
8 bands from random numbers
C
C
Atomic positions and initial magnetic moments
Positions:
0 C 0.000000 0.000000 0.000000 ( 0.0000, 0.0000, 0.0000)
1 C 0.806750 0.806750 0.806750 ( 0.0000, 0.0000, 0.0000)
Unit cell:
periodic x y z points spacing
1. axis: yes 0.000000 1.613500 1.613500 12 0.1553
2. axis: yes 1.613500 0.000000 1.613500 12 0.1553
3. axis: yes 1.613500 1.613500 0.000000 12 0.1553
Lengths: 2.281834 2.281834 2.281834
Angles: 60.000000 60.000000 60.000000
Effective grid spacing dv^(1/3) = 0.1694
iter time total log10-change:
energy eigst dens
iter: 1 12:57:29 -10.023943 +1.54 c
iter: 2 12:57:29 -10.024911 -0.61 c
iter: 3 12:57:29 -10.024916c -2.83 c
iter: 4 12:57:30 -10.024916c -5.07 c
iter: 5 12:57:30 -10.024916c -7.24 c
iter: 6 12:57:31 -10.024916c -9.38c c
Converged after 6 iterations.
Dipole moment: (-1.857210, -1.857210, -1.857210) |e|*Ang
Energy contributions relative to reference atoms: (reference = -2055.256231)
Kinetic: +58.466655
Potential: -41.548711
External: +0.000000
XC: -26.955332
Entropy (-ST): -0.000000
Local: +0.012472
SIC: +0.000000
--------------------------
Free energy: -10.024916
Extrapolated: -10.024916
Showing only first 2 kpts
Kpt Band Eigenvalues Occupancy
0 2 6.44407 2.00000
0 3 6.44414 2.00000
0 4 29.71457 0.00000
0 5 29.71459 0.00000
1 2 6.16314 2.00000
1 3 7.55668 2.00000
1 4 28.81707 0.00000
1 5 30.36033 0.00000
Fermi level: 19.61555
Gap: 4.845 eV
Transition (v -> c):
(s=0, k=59, n=3, [0.00, 0.00, 0.00]) -> (s=0, k=29, n=4, [0.38, 0.00, 0.38])
Direct/indirect transitions are different
Direct gap: 6.434 eV
Transition at: (s=0, k=19, n=3, [0.00, 0.00, 0.00])
Writing to diamond-bands.gpw (mode='all')
[4]:
calc_bands = GPAW(f"{seed}-bands.gpw",)
bs_dft = calc_bands.band_structure()
bs_dft.plot(show=True, emax=40.0)
Timing: incl. excl.
-----------------------------------------------------------
Hamiltonian: 0.019 0.000 0.0% |
Atomic: 0.017 0.000 0.0% |
XC Correction: 0.017 0.017 0.1% |
Calculate atomic Hamiltonians: 0.000 0.000 0.0% |
Communicate: 0.000 0.000 0.0% |
Initialize Hamiltonian: 0.000 0.000 0.0% |
Poisson: 0.000 0.000 0.0% |
XC 3D grid: 0.002 0.002 0.0% |
LCAO initialization: 0.642 0.006 0.0% |
LCAO eigensolver: 0.181 0.001 0.0% |
Calculate projections: 0.002 0.002 0.0% |
DenseAtomicCorrection: 0.006 0.006 0.0% |
Distribute overlap matrix: 0.001 0.001 0.0% |
Orbital Layouts: 0.008 0.008 0.1% |
Potential matrix: 0.151 0.151 1.0% |
Sum over cells: 0.012 0.012 0.1% |
LCAO to grid: 0.186 0.186 1.3% ||
Set positions (LCAO WFS): 0.268 0.036 0.2% |
Basic WFS set positions: 0.000 0.000 0.0% |
Basis functions set positions: 0.000 0.000 0.0% |
P tci: 0.047 0.047 0.3% |
ST tci: 0.174 0.174 1.2% |
mktci: 0.011 0.011 0.1% |
PWDescriptor: 0.002 0.002 0.0% |
SCF-cycle: 2.571 0.009 0.1% |
Davidson: 2.544 0.325 2.2% ||
Apply H: 0.440 0.427 2.9% ||
HMM T: 0.013 0.013 0.1% |
Subspace diag: 0.547 0.009 0.1% |
calc_h_matrix: 0.481 0.038 0.3% |
Apply H: 0.444 0.429 3.0% ||
HMM T: 0.014 0.014 0.1% |
diagonalize: 0.033 0.033 0.2% |
rotate_psi: 0.023 0.023 0.2% |
calc. matrices: 1.058 0.182 1.3% ||
Apply H: 0.876 0.850 5.9% |-|
HMM T: 0.026 0.026 0.2% |
diagonalize: 0.128 0.128 0.9% |
rotate_psi: 0.047 0.047 0.3% |
Orthonormalize: 0.017 0.001 0.0% |
calc_s_matrix: 0.004 0.004 0.0% |
inverse-cholesky: 0.003 0.003 0.0% |
projections: 0.007 0.007 0.0% |
rotate_psi_s: 0.002 0.002 0.0% |
Set symmetry: 0.000 0.000 0.0% |
Other: 11.284 11.284 77.7% |------------------------------|
-----------------------------------------------------------
Total: 14.519 100.0%
Memory usage: 282.95 MiB
Date: Thu Apr 2 12:57:42 2026
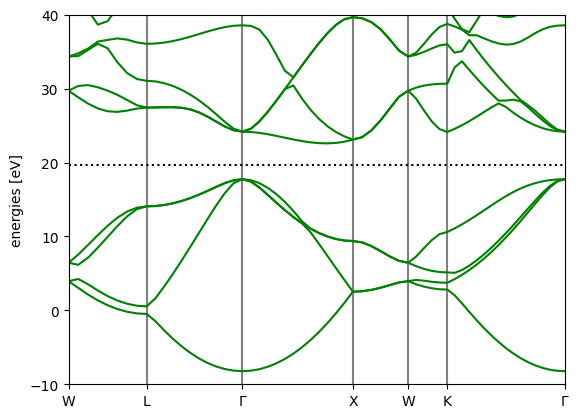
[4]:
<Axes: ylabel='energies [eV]'>
Step 2: Wannierization
In this case we use sp3 projections on each carbon atom.
[20]:
from wannierberri.symmetry.projections import Projection, ProjectionsSet
from wannierberri.w90files import WannierData
calc_scf = GPAW(f'{seed}-scf.gpw', txt=None)
sg = SpaceGroup.from_gpaw(calc_scf)
positions = sg.positions
proj_sp3 = Projection(
position_num=positions,
orbital='sp3',
spacegroup=sg,
rotate_basis=True
)
proj_set = ProjectionsSet(projections=[proj_sp3])
typat used for spacegroup detection (accounting magmom): [6 6]
create the “w90 files”
Here we are NOT using the gpaw-wannier90 interface, and actually not creating the w90 files, but directly access the wavefunctions from the GPAW calculation, use symmetry operations from irrep, and create the necessary data to be used with wannierberri. Those files still retain the same naming convention as the w90 files for consistency, but htey are binary npz files, which are convenient to work with numpy
[21]:
# seedname_ref = os.path.join(path_data, "diamond-irred")
calc_nscf_irred = GPAW(f'{seed}-nscf.gpw', txt=None)
wandata, bandstructure = WannierData.from_gpaw(
calculator=calc_nscf_irred,
spin_channel=0,
projections=proj_set,
select_grid=(4,4,4),
irreducible=True,
files=["amn", "mmn", "eig", "symmetrizer"],
unitary_params=dict(error_threshold=0.1,
warning_threshold=0.01,
nbands_upper_skip=8),
return_bandstructure=True
)
wandata.to_npz(f"{seed}-sp3")
finding num points from 1 projections
finding num points from 1 projections
got irreducible=True, mp_grid=None, seedname=wannier, files=['amn', 'mmn', 'eig', 'symmetrizer'], projections=ProjectionsSet with 8 Wannier functions and 0 free variables
Projection 0.0, 0.0, 0.0:['sp3'] with 8 Wannier functions (8 per spin x1 spins)
on 2 points (4 per site), unk_grid=None, normalize=True
self.irreducible=True
mpgrid = [4 4 4], 8
Shells found with weights [0.52755554] and tolerance 3.861271010427486e-16
Creating amn. Using projections_set
ProjectionsSet with 8 Wannier functions and 0 free variables
Projection 0.0, 0.0, 0.0:['sp3'] with 8 Wannier functions (8 per spin x1 spins)
on 2 points (4 per site)
NK= 64, selected_kpoints = [0 1 2 3 4 5 6 7], kptirr = [0 1 2 3 4 5 6 7]
saving to diamond-sp3.sawf.npz :
saving to diamond-sp3.bkvec.npz :
saving to diamond-sp3.chk.npz :
saving to diamond-sp3.eig.npz :
saving to diamond-sp3.amn.npz :
saving to diamond-sp3.mmn.npz :
[28]:
wandata.wannierise(
froz_min=-10,
froz_max=30,
outer_min=-10,
outer_max=np.inf,
num_iter=100,
conv_tol=1e-10,
print_progress_every=20,
sitesym=True,
localise=True,
)
wandata.chk.to_npz(f"{seed}-sp3.chk.npz")
/home/stepan/github/wannier-berri/wannierberri/wannierisation/wannierizer.py:248: UserWarning: Ray is not initialized, running in serial mode
warnings.warn("Ray is not initialized, running in serial mode")
Warning: symmetrization did not converge in 10 iterations, final changes 0.004946379178734421, 0.0001965289929905878; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.002736053745220717, 2.173914038202943e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
####################################################################################################
starting WFs
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.131975696017 0.131975696017 0.131975696017 | 0.792353234598
0.131975696017 -0.131975696017 -0.131975696017 | 0.792353234598
-0.131975696017 0.131975696017 -0.131975696017 | 0.792353234598
-0.131975696017 -0.131975696017 0.131975696017 | 0.792353234598
0.674774303983 0.938725696017 0.938725696017 | 0.792353234598
0.938725696017 0.938725696017 0.674774303983 | 0.792353234598
0.674774303983 0.674774303983 0.674774303983 | 0.792353234598
0.938725696017 0.674774303983 0.938725696017 | 0.792353234598
----------------------------------------------------------------------------------------------------
3.227000000000 3.227000000000 3.227000000000 | 6.338825876786 <- sum
maximal spread = 0.792353234598
####################################################################################################
####################################################################################################
Iteration 0 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.132815969473 0.132815969473 0.132815969473 | 0.769489943312
0.132815969473 -0.132815969473 -0.132815969473 | 0.769489943312
-0.132815969473 0.132815969473 -0.132815969473 | 0.769489943312
-0.132815969473 -0.132815969473 0.132815969473 | 0.769489943312
0.673934030527 0.939565969473 0.939565969473 | 0.769489943312
0.939565969473 0.939565969473 0.673934030527 | 0.769489943312
0.673934030527 0.673934030527 0.673934030527 | 0.769489943312
0.939565969473 0.673934030527 0.939565969473 | 0.769489943312
----------------------------------------------------------------------------------------------------
3.227000000000 3.227000000000 3.227000000000 | 6.155919546493 <- sum
maximal spread = 0.769489943312
standard deviation = 0.0
####################################################################################################
####################################################################################################
Iteration 20 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.132625367495 0.132625367495 0.132625367495 | 0.680693515432
0.132625367495 -0.132625367495 -0.132625367495 | 0.680693515432
-0.132625367495 0.132625367495 -0.132625367495 | 0.680693515432
-0.132625367495 -0.132625367495 0.132625367495 | 0.680693515432
0.674124632505 0.939375367495 0.939375367495 | 0.680693515432
0.939375367495 0.939375367495 0.674124632505 | 0.680693515432
0.674124632505 0.674124632505 0.674124632505 | 0.680693515432
0.939375367495 0.674124632505 0.939375367495 | 0.680693515432
----------------------------------------------------------------------------------------------------
3.227000000000 3.227000000000 3.227000000000 | 5.445548123453 <- sum
maximal spread = 0.680693515432
standard deviation = 0.0001787542654021851
####################################################################################################
####################################################################################################
Iteration 40 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.132572802445 0.132572802445 0.132572802445 | 0.681815016774
0.132572802445 -0.132572802445 -0.132572802445 | 0.681815016774
-0.132572802445 0.132572802445 -0.132572802445 | 0.681815016774
-0.132572802445 -0.132572802445 0.132572802445 | 0.681815016774
0.674177197555 0.939322802445 0.939322802445 | 0.681815016774
0.939322802445 0.939322802445 0.674177197555 | 0.681815016774
0.674177197555 0.674177197555 0.674177197555 | 0.681815016774
0.939322802445 0.674177197555 0.939322802445 | 0.681815016774
----------------------------------------------------------------------------------------------------
3.227000000000 3.227000000000 3.227000000000 | 5.454520134189 <- sum
maximal spread = 0.681815016774
standard deviation = 6.668154499543905e-06
####################################################################################################
####################################################################################################
Iteration 60 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.132571761646 0.132571761646 0.132571761646 | 0.681853905015
0.132571761646 -0.132571761646 -0.132571761646 | 0.681853905015
-0.132571761646 0.132571761646 -0.132571761646 | 0.681853905015
-0.132571761646 -0.132571761646 0.132571761646 | 0.681853905015
0.674178238354 0.939321761646 0.939321761646 | 0.681853905015
0.939321761646 0.939321761646 0.674178238354 | 0.681853905015
0.674178238354 0.674178238354 0.674178238354 | 0.681853905015
0.939321761646 0.674178238354 0.939321761646 | 0.681853905015
----------------------------------------------------------------------------------------------------
3.227000000000 3.227000000000 3.227000000000 | 5.454831240118 <- sum
maximal spread = 0.681853905015
standard deviation = 2.2059156019413398e-07
####################################################################################################
####################################################################################################
Iteration 80 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.132571730827 0.132571730827 0.132571730827 | 0.681855187267
0.132571730827 -0.132571730827 -0.132571730827 | 0.681855187267
-0.132571730827 0.132571730827 -0.132571730827 | 0.681855187267
-0.132571730827 -0.132571730827 0.132571730827 | 0.681855187267
0.674178269173 0.939321730827 0.939321730827 | 0.681855187267
0.939321730827 0.939321730827 0.674178269173 | 0.681855187267
0.674178269173 0.674178269173 0.674178269173 | 0.681855187267
0.939321730827 0.674178269173 0.939321730827 | 0.681855187267
----------------------------------------------------------------------------------------------------
3.227000000000 3.227000000000 3.227000000000 | 5.454841498134 <- sum
maximal spread = 0.681855187267
standard deviation = 7.24683054708227e-09
####################################################################################################
####################################################################################################
Final state (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.132571729822 0.132571729822 0.132571729822 | 0.681855229094
0.132571729822 -0.132571729822 -0.132571729822 | 0.681855229094
-0.132571729822 0.132571729822 -0.132571729822 | 0.681855229094
-0.132571729822 -0.132571729822 0.132571729822 | 0.681855229094
0.674178270178 0.939321729822 0.939321729822 | 0.681855229094
0.939321729822 0.939321729822 0.674178270178 | 0.681855229094
0.674178270178 0.674178270178 0.674178270178 | 0.681855229094
0.939321729822 0.674178270178 0.939321729822 | 0.681855229094
----------------------------------------------------------------------------------------------------
3.227000000000 3.227000000000 3.227000000000 | 5.454841832754 <- sum
maximal spread = 0.681855229094
standard deviation = 2.81912869566153e-10
####################################################################################################
time for creating wannierizer 0.12383532524108887
time for iterations 3.3949806690216064
time for updating 3.250436305999756
total time for wannierization 3.5369508266448975
saving to wannier.chk :
saving to diamond-sp3.chk.npz :
[28]:
<wannierberri.w90files.chk.CheckPoint at 0x79b3564c52b0>
Create the System object to be used in WannierBerri calculations
It is called “System_w90” class, although it now does not use the w90 wannierization, but the class is the same.
[23]:
from wannierberri import System_R
system = System_R.from_wannierdata(wandata=wandata, berry=True)
irreducible : True, symmetrize set to True
setting Rvec
setting AA..
setting AA - OK
Real-space lattice:
[[0. 1.6135 1.6135]
[1.6135 0. 1.6135]
[1.6135 1.6135 0. ]]
Number of wannier functions: 8
Number of R points: 123
Recommended size of FFT grid [4 4 4]
num_blocks_left = 1, num_blocks_right = 1
number o R-vectors before symmetrization: 123
number o R-vectors after symmetrization: 291
Step 3: Compute the wannierized bandstructure along the high-symmetry path
create the k path for wannierberri
[24]:
from wannierberri.evaluate_k import evaluate_k_path
from wannierberri.grid import Path
kpoints = {
'G': [0.0, 0.0, 0.0],
'X': [0.5, 0.0, 0.5],
'W': [0.5, 0.25, 0.75],
'L': [0.5, 0.5, 0.5],
'K': [0.375, 0.375, 0.75],
}
path_labels = "WLGXWKG"
path = Path.from_nodes(real_lattice=system.real_lattice,
nodes=[kpoints[label] for label in path_labels],
labels=list(path_labels),
length=100
)
bands_wannier= evaluate_k_path(system, path=path) # length~=2pi/dk
Starting run()
Using the follwing calculators :
############################################################
'tabulate' : <wannierberri.calculators.tabulate.TabulatorAll object at 0x79b35627eab0> :
TabulatorAll - a pack of all k-resolved calculators (Tabulators)
Includes the following tabulators :
--------------------------------------------------
"Energy" : <wannierberri.calculators.tabulate.Energy object at 0x79b357207bf0> : calculator not described
--------------------------------------------------
############################################################
Calculation along a path - checking calculators for compatibility
tabulate <wannierberri.calculators.tabulate.TabulatorAll object at 0x79b35627eab0>
All calculators are compatible
Symmetrization switched off for Path
Grid is regular
The set of k points is a Path() with 140 points and labels {0: 'W', 22: 'L', 49: 'G', 80: 'X', 95: 'W', 106: 'K', 139: 'G'}
generating K_list
Done
Done, sum of weights:140.0
############################################################
Iteration 0 out of 0
processing 140 K points : in serial.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 140 K-points on 1 processes: 0.1118 ; per K-point 0.0008 ; proc-sec per K-point 0.0008
time1 = 2.384185791015625e-07
Totally processed 140 K-points
run() finished
/home/stepan/github/wannier-berri/wannierberri/run_grid.py:207: UserWarning: ray package found, but ray is not initialized, running in serial mode
warnings.warn("ray package found, but ray is not initialized, running in serial mode")
/home/stepan/github/wannier-berri/wannierberri/grid/path.py:272: UserWarning: symmetry is not used for a tabulation along path
warnings.warn("symmetry is not used for a tabulation along path")
Plot the bandstructures
[25]:
from matplotlib import pyplot as plt
fig, ax = plt.subplots(figsize=(8,6))
bs_dft.plot(show=False, emax=40.0, ax=ax, label="DFT")
bands_wannier.plot_path_fat(path=path,
label="wannierised sp3",
# linecolor="orange",
axes=ax,
close_fig=False,
show_fig=False,
kwargs_line=dict(linestyle='--', lw=1.0),
)
plt.show()
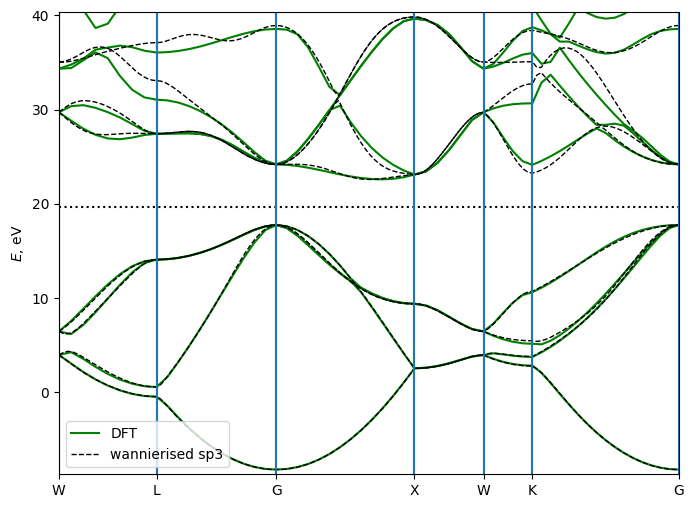
One can see that the wannierized bandstructure (dashed lines) well reproduces the DFT bandstructure (solid lines) in the energy window of interest. Note, that we started from a very coarse DFT grid of 4,4,4 k-points, and still obtained a good agreement after wannierization. You may try to increase the grid on the nscf step, and see how it improves the agreement further.
Bonding and antibonding states
Another option for projections is to use bonding and antibonding states. For that we place the wannier centers on the centers of the bonds, and use s-type projections (or bonding and pz-type for antibonding states, with the “z-axis” along the bond direction [111]. Note, that we specify only one bond position, the other 3 are generated by symmetry operations, and their “zaxis” are also rotated accordingly.
Bonding states describe the valence bands, while antibonding states describe the conduction bands, so they can be wannierised separately.
[32]:
proj_bonding = Projection(
position_num=[1/8, 1/8, 1/8],
orbital='s',
spacegroup=sg,
rotate_basis=True
)
proj_antibonding = Projection(
position_num=[1/8, 1/8, 1/8],
orbital='pz',
zaxis=[1,1,1],
rotate_basis=True,
spacegroup=sg)
proj_set_bonding = ProjectionsSet(projections=[proj_bonding, proj_antibonding])
wandata_bonding = WannierData.from_npz(f"{seed}-sp3", files=["mmn", "symmetrizer", "eig"])
# seedname_ref = os.path.join(path_data, "diamond-irred")
wandata_bonding.set_projections(projections=proj_set_bonding, bandstructure=bandstructure)
wandata_bonding.wannierise(
init="amn",
froz_min=-10,
froz_max=30,
num_iter=1000,
conv_tol=1e-10,
print_progress_every=20,
sitesym=True,
localise=True
)
wandata_bonding.to_npz(f"{seed}-bonding")
system_bonding = System_R.from_wannierdata(wandata=wandata, berry=True)
bands_wannier_bonding = evaluate_k_path(system_bonding,
path=path,
return_path=False)
files = ['mmn', 'symmetrizer', 'eig', 'bkvec']
Trying to read file mmn from npz diamond-sp3.mmn.npz
setting file mmn from npz diamond-sp3.mmn.npz as <wannierberri.w90files.mmn.MMN object at 0x79b3553e8620>
Trying to read file symmetrizer from npz diamond-sp3.symmetrizer.npz
setting file symmetrizer from npz diamond-sp3.sawf.npz as <wannierberri.symmetry.sawf.SymmetrizerSAWF object at 0x79b3553eb2f0>
Trying to read file eig from npz diamond-sp3.eig.npz
setting file eig from npz diamond-sp3.eig.npz as <wannierberri.w90files.eig.EIG object at 0x79b3559a2b70>
Trying to read file bkvec from npz diamond-sp3.bkvec.npz
setting file bkvec from npz diamond-sp3.bkvec.npz as <wannierberri.w90files.bkvectors.BKVectors object at 0x79b356a657c0>
/home/stepan/github/wannier-berri/wannierberri/w90files/wandata.py:405: UserWarning: file mmn cntains 8 k-points less than NK (64) , so we assume the files contain only on irreducible k-points
warnings.warn(f"file {f} cntains {nkeys} k-points less than NK ({NK}) , "
/home/stepan/github/wannier-berri/wannierberri/w90files/wandata.py:405: UserWarning: file eig cntains 8 k-points less than NK (64) , so we assume the files contain only on irreducible k-points
warnings.warn(f"file {f} cntains {nkeys} k-points less than NK ({NK}) , "
/home/stepan/github/wannier-berri/wannierberri/w90files/wandata.py:405: UserWarning: file bkvec cntains 8 k-points less than NK (64) , so we assume the files contain only on irreducible k-points
warnings.warn(f"file {f} cntains {nkeys} k-points less than NK ({NK}) , "
Setting AMN with kwargs={'kptirr': array([0, 1, 2, 3, 4, 5, 6, 7]), 'NK': array(64), 'selected_kpoints': array([0, 1, 2, 3, 4, 5, 6, 7])}
finding num points from 2 projections
finding num points from 2 projections
Creating amn. Using projections_set
ProjectionsSet with 8 Wannier functions and 0 free variables
Projection 0.125, 0.125, 0.125:['s'] with 4 Wannier functions (4 per spin x1 spins)
on 4 points (1 per site)
Projection 0.125, 0.125, 0.125:['pz'] with 4 Wannier functions (4 per spin x1 spins)
on 4 points (1 per site)
AMN file created with NK=64
Warning: symmetrization did not converge in 10 iterations, final changes 0.009941136893744167, 0.0005654612548233089; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.0025048962377636737, 1.3670316302605833e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
/home/stepan/github/wannier-berri/wannierberri/wannierisation/wannierizer.py:248: UserWarning: Ray is not initialized, running in serial mode
warnings.warn("Ray is not initialized, running in serial mode")
####################################################################################################
starting WFs
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.403375000000 0.403375000000 0.403375000000 | 0.717340997200
0.403375000000 1.210125000000 1.210125000000 | 0.717340997200
1.210125000000 1.210125000000 0.403375000000 | 0.717340997200
1.210125000000 0.403375000000 1.210125000000 | 0.717340997200
0.403375000000 0.403375000000 0.403375000000 | 1.748464055419
0.403375000000 1.210125000000 1.210125000000 | 1.748464055419
1.210125000000 1.210125000000 0.403375000000 | 1.748464055419
1.210125000000 0.403375000000 1.210125000000 | 1.748464055419
----------------------------------------------------------------------------------------------------
6.454000000000 6.454000000000 6.454000000000 | 9.863220210474 <- sum
maximal spread = 1.748464055419
####################################################################################################
####################################################################################################
Iteration 0 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.403375000000 0.403375000000 0.403375000000 | 0.693972703324
0.403375000000 1.210125000000 1.210125000000 | 0.693972703324
1.210125000000 1.210125000000 0.403375000000 | 0.693972703324
1.210125000000 0.403375000000 1.210125000000 | 0.693972703324
0.403375000000 0.403375000000 0.403375000000 | 1.466943839620
0.403375000000 1.210125000000 1.210125000000 | 1.466943839620
1.210125000000 1.210125000000 0.403375000000 | 1.466943839620
1.210125000000 0.403375000000 1.210125000000 | 1.466943839620
----------------------------------------------------------------------------------------------------
6.454000000000 6.454000000000 6.454000000000 | 8.643666171776 <- sum
maximal spread = 1.466943839620
standard deviation = 0.0
####################################################################################################
####################################################################################################
Iteration 20 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.403375000000 0.403375000000 0.403375000000 | 0.529120846132
0.403375000000 1.210125000000 1.210125000000 | 0.529120846132
1.210125000000 1.210125000000 0.403375000000 | 0.529120846132
1.210125000000 0.403375000000 1.210125000000 | 0.529120846132
0.403375000000 0.403375000000 0.403375000000 | 1.148138059064
0.403375000000 1.210125000000 1.210125000000 | 1.148138059064
1.210125000000 1.210125000000 0.403375000000 | 1.148138059064
1.210125000000 0.403375000000 1.210125000000 | 1.148138059064
----------------------------------------------------------------------------------------------------
6.454000000000 6.454000000000 6.454000000000 | 6.709035620784 <- sum
maximal spread = 1.148138059064
standard deviation = 0.0007237403300503724
####################################################################################################
####################################################################################################
Iteration 40 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.403375000000 0.403375000000 0.403375000000 | 0.527074033341
0.403375000000 1.210125000000 1.210125000000 | 0.527074033341
1.210125000000 1.210125000000 0.403375000000 | 0.527074033341
1.210125000000 0.403375000000 1.210125000000 | 0.527074033341
0.403375000000 0.403375000000 0.403375000000 | 1.152355993009
0.403375000000 1.210125000000 1.210125000000 | 1.152355993009
1.210125000000 1.210125000000 0.403375000000 | 1.152355993009
1.210125000000 0.403375000000 1.210125000000 | 1.152355993009
----------------------------------------------------------------------------------------------------
6.454000000000 6.454000000000 6.454000000000 | 6.717720105401 <- sum
maximal spread = 1.152355993009
standard deviation = 2.317626220501394e-05
####################################################################################################
####################################################################################################
Iteration 60 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.403375000000 0.403375000000 0.403375000000 | 0.527017375425
0.403375000000 1.210125000000 1.210125000000 | 0.527017375425
1.210125000000 1.210125000000 0.403375000000 | 0.527017375425
1.210125000000 0.403375000000 1.210125000000 | 0.527017375425
0.403375000000 0.403375000000 0.403375000000 | 1.152488940766
0.403375000000 1.210125000000 1.210125000000 | 1.152488940766
1.210125000000 1.210125000000 0.403375000000 | 1.152488940766
1.210125000000 0.403375000000 1.210125000000 | 1.152488940766
----------------------------------------------------------------------------------------------------
6.454000000000 6.454000000000 6.454000000000 | 6.718025264764 <- sum
maximal spread = 1.152488940766
standard deviation = 7.351141257177589e-07
####################################################################################################
####################################################################################################
Iteration 80 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.403375000000 0.403375000000 0.403375000000 | 0.527015636841
0.403375000000 1.210125000000 1.210125000000 | 0.527015636841
1.210125000000 1.210125000000 0.403375000000 | 0.527015636841
1.210125000000 0.403375000000 1.210125000000 | 0.527015636841
0.403375000000 0.403375000000 0.403375000000 | 1.152493186086
0.403375000000 1.210125000000 1.210125000000 | 1.152493186086
1.210125000000 1.210125000000 0.403375000000 | 1.152493186086
1.210125000000 0.403375000000 1.210125000000 | 1.152493186086
----------------------------------------------------------------------------------------------------
6.454000000000 6.454000000000 6.454000000000 | 6.718035291709 <- sum
maximal spread = 1.152493186086
standard deviation = 2.3749610336208095e-08
####################################################################################################
####################################################################################################
Iteration 100 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.403375000000 0.403375000000 0.403375000000 | 0.527015581338
0.403375000000 1.210125000000 1.210125000000 | 0.527015581338
1.210125000000 1.210125000000 0.403375000000 | 0.527015581338
1.210125000000 0.403375000000 1.210125000000 | 0.527015581338
0.403375000000 0.403375000000 0.403375000000 | 1.152493323678
0.403375000000 1.210125000000 1.210125000000 | 1.152493323678
1.210125000000 1.210125000000 0.403375000000 | 1.152493323678
1.210125000000 0.403375000000 1.210125000000 | 1.152493323678
----------------------------------------------------------------------------------------------------
6.454000000000 6.454000000000 6.454000000000 | 6.718035620064 <- sum
maximal spread = 1.152493323678
standard deviation = 7.736100090873036e-10
####################################################################################################
Converged after 112 iterations
####################################################################################################
Final state (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.403375000000 0.403375000000 0.403375000000 | 0.527015579715
0.403375000000 1.210125000000 1.210125000000 | 0.527015579715
1.210125000000 1.210125000000 0.403375000000 | 0.527015579715
1.210125000000 0.403375000000 1.210125000000 | 0.527015579715
0.403375000000 0.403375000000 0.403375000000 | 1.152493327722
0.403375000000 1.210125000000 1.210125000000 | 1.152493327722
1.210125000000 1.210125000000 0.403375000000 | 1.152493327722
1.210125000000 0.403375000000 1.210125000000 | 1.152493327722
----------------------------------------------------------------------------------------------------
6.454000000000 6.454000000000 6.454000000000 | 6.718035629748 <- sum
maximal spread = 1.152493327722
standard deviation = 9.931148324964522e-11
####################################################################################################
time for creating wannierizer 0.1302335262298584
time for iterations 6.261308670043945
time for updating 6.10145902633667
total time for wannierization 6.435135126113892
saving to diamond-sp3.chk :
saving to diamond-bonding.mmn.npz :
saving to diamond-bonding.sawf.npz :
saving to diamond-bonding.eig.npz :
saving to diamond-bonding.bkvec.npz :
saving to diamond-bonding.amn.npz :
saving to diamond-bonding.chk.npz :
irreducible : True, symmetrize set to True
setting Rvec
setting AA..
setting AA - OK
Real-space lattice:
[[0. 1.6135 1.6135]
[1.6135 0. 1.6135]
[1.6135 1.6135 0. ]]
Number of wannier functions: 8
Number of R points: 123
Recommended size of FFT grid [4 4 4]
num_blocks_left = 1, num_blocks_right = 1
number o R-vectors before symmetrization: 123
number o R-vectors after symmetrization: 291
Starting run()
Using the follwing calculators :
############################################################
'tabulate' : <wannierberri.calculators.tabulate.TabulatorAll object at 0x79b356e40740> :
TabulatorAll - a pack of all k-resolved calculators (Tabulators)
Includes the following tabulators :
--------------------------------------------------
"Energy" : <wannierberri.calculators.tabulate.Energy object at 0x79b3559169c0> : calculator not described
--------------------------------------------------
############################################################
Calculation along a path - checking calculators for compatibility
tabulate <wannierberri.calculators.tabulate.TabulatorAll object at 0x79b356e40740>
All calculators are compatible
Symmetrization switched off for Path
Grid is regular
The set of k points is a Path() with 140 points and labels {0: 'W', 22: 'L', 49: 'G', 80: 'X', 95: 'W', 106: 'K', 139: 'G'}
generating K_list
Done
Done, sum of weights:140.0
############################################################
Iteration 0 out of 0
processing 140 K points : in serial.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 140 K-points on 1 processes: 0.1136 ; per K-point 0.0008 ; proc-sec per K-point 0.0008
time1 = 4.76837158203125e-07
Totally processed 140 K-points
run() finished
/home/stepan/github/wannier-berri/wannierberri/run_grid.py:207: UserWarning: ray package found, but ray is not initialized, running in serial mode
warnings.warn("ray package found, but ray is not initialized, running in serial mode")
/home/stepan/github/wannier-berri/wannierberri/grid/path.py:272: UserWarning: symmetry is not used for a tabulation along path
warnings.warn("symmetry is not used for a tabulation along path")
[33]:
from matplotlib import pyplot as plt
fig, ax = plt.subplots(figsize=(8,6))
bs_dft.plot(filename='bandstructure.png', show=False, emax=40.0, ax=ax, label="DFT")
bands_wannier_bonding.plot_path_fat(path=path,
label="wannierised-bonding",
# linecolor="orange",
axes=ax,
close_fig=False,
show_fig=False,
kwargs_line=dict(linestyle='--', lw=1.0),
)
plt.show()
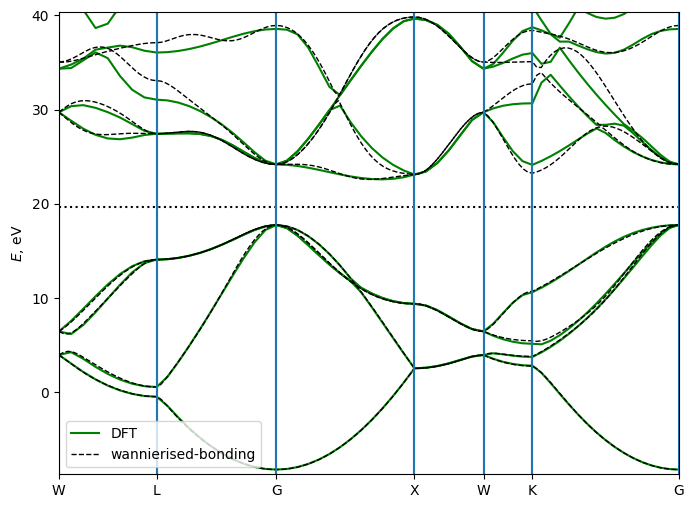
Save the system objects for later use
[35]:
system_bonding.save_npz("system-diamond-bonding")
system.save_npz("system-diamond-sp3")
# load the system later
from wannierberri.system.system_R import System_R
system_loaded = System_R.from_npz("system-diamond-sp3")
saving system of class System_R to system-diamond-bonding
properties: ['num_wann', 'real_lattice', 'iRvec', 'periodic', 'is_phonon', 'wannier_centers_cart', 'pointgroup']
saving num_wann
saving num_wann to system-diamond-bonding/num_wann.npz
- Ok!
saving real_lattice
saving real_lattice to system-diamond-bonding/real_lattice.npz
- Ok!
saving iRvec
saving iRvec to system-diamond-bonding/iRvec.npz
- Ok!
saving periodic
saving periodic to system-diamond-bonding/periodic.npz
- Ok!
saving is_phonon
saving is_phonon to system-diamond-bonding/is_phonon.npz
- Ok!
saving wannier_centers_cart
saving wannier_centers_cart to system-diamond-bonding/wannier_centers_cart.npz
- Ok!
saving pointgroup
saving pointgroup to system-diamond-bonding/pointgroup.npz
- Ok!
saving Ham - Ok!
saving AA - Ok!
saving system of class System_R to system-diamond-sp3
properties: ['num_wann', 'real_lattice', 'iRvec', 'periodic', 'is_phonon', 'wannier_centers_cart', 'pointgroup']
saving num_wann
saving num_wann to system-diamond-sp3/num_wann.npz
- Ok!
saving real_lattice
saving real_lattice to system-diamond-sp3/real_lattice.npz
- Ok!
saving iRvec
saving iRvec to system-diamond-sp3/iRvec.npz
- Ok!
saving periodic
saving periodic to system-diamond-sp3/periodic.npz
- Ok!
saving is_phonon
saving is_phonon to system-diamond-sp3/is_phonon.npz
- Ok!
saving wannier_centers_cart
saving wannier_centers_cart to system-diamond-sp3/wannier_centers_cart.npz
- Ok!
saving pointgroup
saving pointgroup to system-diamond-sp3/pointgroup.npz
- Ok!
saving Ham - Ok!
saving AA - Ok!
loading real_lattice - Ok!
loading wannier_centers_cart - Ok!
loading pointgroup - Ok!
loading iRvec - Ok!
loading periodic - Ok!
loading is_phonon - Ok!
loading atom_labels - Ok!
loading num_wann - Ok!
loading positions - Ok!
loading R_matrix AA - Ok!
loading R_matrix Ham - Ok!
[ ]: