GPAW + WannierBerri: Altermagnet MnTe
It is assumed that the reader is familiar with the previous tutorial on bcc Fe with GPAW. In this tutorial we will show a special case, the crystal structure is anti-ferromagnetic. MnTe has two Mn atoms per unit cell with opposite directions of magnetic moments. As a result in spin-up states, the Mn1 d-states are occupied, while the Mn2 d-states are empty. In the spin-down channel the situation is reversed. Thus, to wannierise only the valence bands, one may want to introduce different projections for the two spin channels.
[1]:
import numpy as np
from ase import Atoms
from gpaw import GPAW, PW, MixerSum
import ray
ray.init(num_cpus=16)
seed = "MnTe"
a = 4.134
c = 6.652
lattice = a * np.array([[1, 0, 0], [-1 / 2, np.sqrt(3) / 2, 0], [0, 0, c / a]])
positions = np.array(
[
[0, 0, 0],
[0, 0, 1 / 2],
[1 / 3, 2 / 3, 1 / 4],
[2 / 3, 1 / 3, 3 / 4],
]
)
/home/stepan/github/WannierBerri-tutorial/.conda/lib/python3.12/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
2026-04-02 16:55:25,442 INFO util.py:154 -- Missing packages: ['ipywidgets']. Run `pip install -U ipywidgets`, then restart the notebook server for rich notebook output.
2026-04-02 16:55:27,612 INFO worker.py:1918 -- Started a local Ray instance. View the dashboard at http://127.0.0.1:8265
Step 1: Gpaw calculation
loading the GPAW calculators
If you want to skip the GPAW calculation, you can just load the pre-calculated calculators (not included in the repository due to size, but if you have run gpaw once, you can find them in the gpaw folder).
[2]:
calc_scf = GPAW(f"gpaw/{seed}-scf.gpw")
calc_nscf = GPAW(f"gpaw/{seed}-nscf-irred-664.gpw")
calc_bands = GPAW(f"gpaw/{seed}-bands-path.gpw")
self-consistent calculation
[ ]:
a = 4.134
c = 6.652
lattice = a * np.array([[1, 0, 0], [-1 / 2, np.sqrt(3) / 2, 0], [0, 0, c / a]])
positions = np.array(
[
[0, 0, 0],
[0, 0, 1 / 2],
[1 / 3, 2 / 3, 1 / 4],
[2 / 3, 1 / 3, 3 / 4],
]
)
atoms = Atoms("Mn2Te2",
cell=lattice,
scaled_positions=positions,
pbc=[1, 1, 1])
m = 4.7
magmoms = np.zeros(4)
magmoms[0] = +m
magmoms[1] = -m
atoms.set_initial_magnetic_moments(magmoms)
calc = GPAW(
mode=PW(600),
xc="PBE",
kpts={"size": [6, 6, 4], "gamma": True},
convergence={"density": 1e-6},
mixer=MixerSum(0.25, 8, 100),
setups={"Mn": ":d,4.0"},
txt="rpaw/MnTe_scf.txt"
)
atoms.calc = calc
atoms.get_potential_energy()
calc.write(f"gpaw/{seed}-scf.gpw", mode="all")
Non-self-consistent calculation in the irreducible Brillouin zone
Now we get the list of irreducible k-points (using irrep.SpaceGroup object) and perform a non-self-consistent calculation only on these k-points. Note, that here we use all symmetries of the crystal structure, even those which are broken by the magnetization direction, because they are still symmetries for each spin channel separately (including the time-reversal symmetry).
[ ]:
from irrep.spacegroup import SpaceGroup
sg = SpaceGroup.from_gpaw(calc)
kpoints_irred = sg.get_irreducible_kpoints_grid([6, 6, 4])
calc_nscf = calc.fixed_density(
kpts=kpoints_irred,
symmetry={'symmorphic': False},
nbands=60,
convergence={'bands': 40},
txt=f'gpaw/{seed}-nscf.txt')
calc_nscf.write(f'gpaw/{seed}-nscf-irred-664.gpw', mode='all')
typat used for spacegroup detection (accounting magmoms): [26]
Compute the dft bandstructure along a high-symmetry path
This is done to compare with the wannierized bandstructure later.
[ ]:
from wannierberri.grid import Path
# recip_lattice = 2*np.pi * np.linalg.inv(lattice).T
kz = 0.35 / (2*np.pi/c)
path = Path(lattice,
nodes=[
[2 / 3, -1 / 3, 0],
[0, 0, 0],
[-2 / 3, 1 / 3, 0],
None,
[-0.5, 0, kz],
[0, 0, kz],
[0.5, 0, kz],
],
labels=[r"${\rm K}\leftarrow$",
r"$\Gamma$",
r"$\rightarrow{\rm K}$",
r"$\overline{\rm M}\leftarrow$",
r"$\overline{\Gamma}$",
r"$\rightarrow\overline{\rm M}$"],
length=100) # length [ Ang] ~= 2*pi/dk
kpoints = path.K_list
print (kpoints.shape)
print (kpoints)
calc_bands = calc_scf.fixed_density(
nbands=30,
symmetry='off',
random=True,
kpts=kpoints,
txt=f'gpaw/{seed}-bands-path.txt',
convergence={'bands': 24})
calc_bands.write(f'gpaw/{seed}-bands-path.gpw', mode='all')
(61, 3)
[[ 0.66666667 -0.33333333 0. ]
[ 0.625 -0.3125 0. ]
[ 0.58333333 -0.29166667 0. ]
[ 0.54166667 -0.27083333 0. ]
[ 0.5 -0.25 0. ]
[ 0.45833333 -0.22916667 0. ]
[ 0.41666667 -0.20833333 0. ]
[ 0.375 -0.1875 0. ]
[ 0.33333333 -0.16666667 0. ]
[ 0.29166667 -0.14583333 0. ]
[ 0.25 -0.125 0. ]
[ 0.20833333 -0.10416667 0. ]
[ 0.16666667 -0.08333333 0. ]
[ 0.125 -0.0625 0. ]
[ 0.08333333 -0.04166667 0. ]
[ 0.04166667 -0.02083333 0. ]
[ 0. 0. 0. ]
[-0.04166667 0.02083333 0. ]
[-0.08333333 0.04166667 0. ]
[-0.125 0.0625 0. ]
[-0.16666667 0.08333333 0. ]
[-0.20833333 0.10416667 0. ]
[-0.25 0.125 0. ]
[-0.29166667 0.14583333 0. ]
[-0.33333333 0.16666667 0. ]
[-0.375 0.1875 0. ]
[-0.41666667 0.20833333 0. ]
[-0.45833333 0.22916667 0. ]
[-0.5 0.25 0. ]
[-0.54166667 0.27083333 0. ]
[-0.58333333 0.29166667 0. ]
[-0.625 0.3125 0. ]
[-0.5 0. 0.37054454]
[-0.46428571 0. 0.37054454]
[-0.42857143 0. 0.37054454]
[-0.39285714 0. 0.37054454]
[-0.35714286 0. 0.37054454]
[-0.32142857 0. 0.37054454]
[-0.28571429 0. 0.37054454]
[-0.25 0. 0.37054454]
[-0.21428571 0. 0.37054454]
[-0.17857143 0. 0.37054454]
[-0.14285714 0. 0.37054454]
[-0.10714286 0. 0.37054454]
[-0.07142857 0. 0.37054454]
[-0.03571429 0. 0.37054454]
[ 0. 0. 0.37054454]
[ 0.03571429 0. 0.37054454]
[ 0.07142857 0. 0.37054454]
[ 0.10714286 0. 0.37054454]
[ 0.14285714 0. 0.37054454]
[ 0.17857143 0. 0.37054454]
[ 0.21428571 0. 0.37054454]
[ 0.25 0. 0.37054454]
[ 0.28571429 0. 0.37054454]
[ 0.32142857 0. 0.37054454]
[ 0.35714286 0. 0.37054454]
[ 0.39285714 0. 0.37054454]
[ 0.42857143 0. 0.37054454]
[ 0.46428571 0. 0.37054454]
[ 0.5 0. 0.37054454]]
[3]:
bs_dft = calc_bands.band_structure()
bs_dft.plot(show=True, emin=-8,emax=15.0)
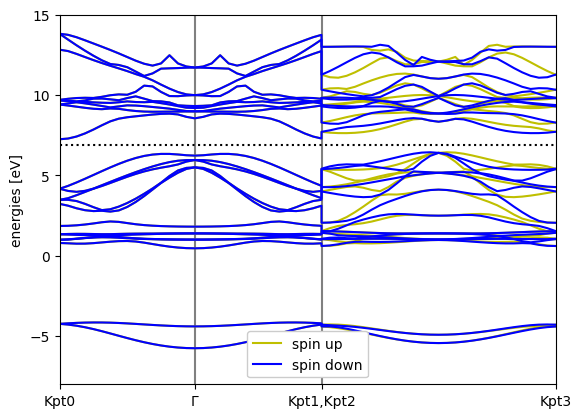
[3]:
<Axes: ylabel='energies [eV]'>
Step 2: Wannierization
In this case we use sp3d2 and t2g projections on each iron atom. One can also use the conventional s,p,d orbitals.
[4]:
from irrep.spacegroup import SpaceGroup
from wannierberri.symmetry.projections import Projection, ProjectionsSet
sg = SpaceGroup.from_gpaw(calc_nscf)
positions_Mn = [[0, 0, 0],
[0, 0, 1 / 2]]
positions_Te = [[1 / 3, 2 / 3, 1 / 4],
[2 / 3, 1 / 3, 3 / 4]]
proj_Mn1_d = Projection(position_num=positions_Mn[0], orbital='d', spacegroup=sg)
proj_Mn2_d = Projection(position_num=positions_Mn[1], orbital='d', spacegroup=sg)
proj_Te_sp2 = Projection(position_num=positions_Te, orbital='sp2', spacegroup=sg, xaxis=[0, -1, 0], rotate_basis=True)
proj_Te_pz = Projection(position_num=positions_Te, orbital='pz', spacegroup=sg)
proj_set_up = ProjectionsSet([proj_Mn1_d, proj_Te_sp2, proj_Te_pz])
proj_set_down = ProjectionsSet([proj_Mn2_d, proj_Te_sp2, proj_Te_pz])
typat used for spacegroup detection (accounting magmom): [25003, 25001, 52002, 52002]
create the “w90 files”
Here we are NOT using the gpaw-wannier90 interface, and actually not creating the w90 files, but directly access the wavefunctions from the GPAW calculation, use symmetry operations from irrep, and create the necessary data to be used with wannierberri. Those files still retain the same naming convention as the w90 files for consistency, but htey are binary npz files, which are convenient to work with numpy
After creating those data, it is written to the disk, so that one can later load it. But be careful - if you change anything (e.g. the projections) - do not forget to remove those files, otherwise you will load the old data! For that reason, here we set ‘read_npz_list=[]’ to force the creation of new data.
[10]:
from wannierberri.w90files.w90data_soc import Wannier90dataSOC
wandata = Wannier90dataSOC.from_gpaw(
seedname="wannier_soc",
calculator=calc_nscf,
projections_up=proj_set_up,
projections_down=proj_set_down,
mp_grid=(6, 6, 4),
spacegroup=sg
)
wandata.to_npz("wannier_soc")
got irreducible=None, mp_grid=(6, 6, 4), seedname=wannier_soc-spin-0, files=['mmn', 'eig', 'amn', 'symmetrizer'], projections=ProjectionsSet with 13 Wannier functions and 0 free variables
Projection 0, 0, 0:['d'] with 5 Wannier functions (5 per spin x1 spins)
on 1 points (5 per site)
Projection 0.3333333333333333, 0.6666666666666666, 0.25:['sp2'] with 6 Wannier functions (6 per spin x1 spins)
on 2 points (3 per site)
Projection 0.3333333333333333, 0.6666666666666666, 0.25:['pz'] with 2 Wannier functions (2 per spin x1 spins)
on 2 points (1 per site), unk_grid=None, normalize=True
mpgrid = [6 6 4], 24
detected grid=(np.int64(6), np.int64(6), np.int64(4)), selected_kpoints=[ 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23]
self.irreducible=True
mpgrid = [6 6 4], 24
/home/stepan/github/wannier-berri/wannierberri/w90files/wandata.py:128: UserWarning: detected grid (np.int64(6), np.int64(6), np.int64(4)) od 144 kpoints, but only 24 kpoints are available.assuming that only irreducible kpoints are needed.
warnings.warn(f"detected grid {grid} od {np.prod(grid)} kpoints, "
Shells found with weights [8.96674318 3.89604278] and tolerance 3.147886549272457e-16
Creating amn. Using projections_set
ProjectionsSet with 13 Wannier functions and 0 free variables
Projection 0, 0, 0:['d'] with 5 Wannier functions (5 per spin x1 spins)
on 1 points (5 per site)
Projection 0.3333333333333333, 0.6666666666666666, 0.25:['sp2'] with 6 Wannier functions (6 per spin x1 spins)
on 2 points (3 per site)
Projection 0.3333333333333333, 0.6666666666666666, 0.25:['pz'] with 2 Wannier functions (2 per spin x1 spins)
on 2 points (1 per site)
NK= 144, selected_kpoints = [ 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23], kptirr = [ 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23]
got irreducible=None, mp_grid=(6, 6, 4), seedname=wannier_soc-spin-1, files=['mmn', 'eig', 'amn', 'symmetrizer'], projections=ProjectionsSet with 13 Wannier functions and 0 free variables
Projection 0.0, 0.0, 0.5:['d'] with 5 Wannier functions (5 per spin x1 spins)
on 1 points (5 per site)
Projection 0.3333333333333333, 0.6666666666666666, 0.25:['sp2'] with 6 Wannier functions (6 per spin x1 spins)
on 2 points (3 per site)
Projection 0.3333333333333333, 0.6666666666666666, 0.25:['pz'] with 2 Wannier functions (2 per spin x1 spins)
on 2 points (1 per site), unk_grid=None, normalize=True
mpgrid = [6 6 4], 24
detected grid=(np.int64(6), np.int64(6), np.int64(4)), selected_kpoints=[ 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23]
self.irreducible=True
mpgrid = [6 6 4], 24
Creating amn. Using projections_set
ProjectionsSet with 13 Wannier functions and 0 free variables
Projection 0.0, 0.0, 0.5:['d'] with 5 Wannier functions (5 per spin x1 spins)
on 1 points (5 per site)
Projection 0.3333333333333333, 0.6666666666666666, 0.25:['sp2'] with 6 Wannier functions (6 per spin x1 spins)
on 2 points (3 per site)
Projection 0.3333333333333333, 0.6666666666666666, 0.25:['pz'] with 2 Wannier functions (2 per spin x1 spins)
on 2 points (1 per site)
NK= 144, selected_kpoints = [ 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23], kptirr = [ 0 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23]
number of bands = 60
saving to wannier_soc.soc.npz :
saving to wannier_soc-spin-0.sawf.npz :
saving to wannier_soc-spin-0.bkvec.npz :
saving to wannier_soc-spin-0.chk.npz :
saving to wannier_soc-spin-0.eig.npz :
saving to wannier_soc-spin-0.amn.npz :
saving to wannier_soc-spin-0.mmn.npz :
saving to wannier_soc-spin-1.sawf.npz :
saving to wannier_soc-spin-1.bkvec.npz :
saving to wannier_soc-spin-1.chk.npz :
saving to wannier_soc-spin-1.eig.npz :
saving to wannier_soc-spin-1.amn.npz :
saving to wannier_soc-spin-1.mmn.npz :
Do the wannierization
at this step we also specify the frozen window, number of iterations, and whether to use site symmetry and localization. (when starting from irreducible BZ, site-symmetry is required anyway)
We also may use the outer window via w90data.select_bands if needed
In this case, both spin channels will be wannierised separately, starting from the same projections.There is also an option to specify different projections for different spin channels if needed. (See MnTe tutorial)
[11]:
wandata.wannierise(
froz_min=-10,
froz_max=7,
outer_min=-10,
outer_max=np.inf,
num_iter=300,
print_progress_every=50,
sitesym=True,
localise=True,
)
Warning: symmetrization did not converge in 10 iterations, final changes 0.006164440129185927, 5.4168095907182255e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.003664581914962914, 2.1742096396759088e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=1, kpt=1, [0. 0. 0.25] with 12 symmetries, max error in included blocks: 4.625414467873572e-06 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
Warning: symmetrization did not converge in 10 iterations, final changes 0.006561222140921279, 0.00110052061772105; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.00013001646791354848, 4.238676422731344e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.005319834071425336, 0.0005067463655086965; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 37 [55:56] is 4.625414467873572e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
Warning: symmetrization did not converge in 10 iterations, final changes 6.502544588495062e-05, 5.5252166721961275e-06; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 1.5830410170095906e-05, 3.0803098126613112e-06; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=8, kpt=8, [0. 0.33333333 0.25 ] with 4 symmetries, max error in included blocks: 1.4171197572519495e-06 ; excluded bands are [57 58 59] out of 60 total bands (accuracy threshold 1e-06)
Warning: symmetrization did not converge in 10 iterations, final changes 0.017230485375504336, 0.002215824648506568; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 3.825254840807255e-06, 8.767160718800709e-07; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.0007713032754051138, 0.00018186509049658006; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=11, kpt=11, [0. 0.5 0. ] with 8 symmetries, max error in included blocks: 5.099233808554998e-06 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 50 [54:55] is 1.4171197572519495e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 51 [55:56] is 5.099233808554998e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
Warning: symmetrization did not converge in 10 iterations, final changes 4.390042422705717e-05, 1.4019502096933555e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 1.0351637131574923e-06, 2.3291400492822527e-07; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 1.948033100268342e-06, 2.16411807580461e-07; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.004643632672913008, 0.00020597647291971025; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=21, kpt=21, [0.33333333 0.33333333 0. ] with 12 symmetries, max error in included blocks: 0.0017048129958950444 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
Warning: symmetrization did not converge in 10 iterations, final changes 0.0034454820427515425, 2.2019491002206135e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=22, kpt=22, [0.33333333 0.33333333 0.25 ] with 6 symmetries, max error in included blocks: 1.2577670093556239e-05 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 13 [22:24] is 1.8763315621462945e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 32 [51:53] is 2.5189851632702666e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 33 [53:54] is 0.0009077955136227982, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 34 [54:56] is 0.0017048129958950444, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 15 [25:27] is 1.2577670093556239e-05, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 34 [55:56] is 1.5331304361280537e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
####################################################################################################
starting WFs
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 0.000000000000 | 0.484489492172
0.000000000000 0.000000000000 0.000000000000 | 0.473508056635
0.000000000000 0.000000000000 0.000000000000 | 0.473508056635
0.000000000000 0.000000000000 0.000000000000 | 0.457939241268
0.000000000000 0.000000000000 0.000000000000 | 0.457939241268
0.715030119727 2.799588844933 1.678042683692 | 2.215017821643
-0.715030119727 2.799588844933 1.678042683692 | 2.215017821643
0.000000000000 1.561120348624 1.678042683692 | 2.215017821643
1.351969880273 0.780560174312 4.973957316308 | 2.215017821643
2.782030119727 0.780560174312 4.973957316308 | 2.215017821643
2.067000000000 2.019028670621 4.973957316308 | 2.215017821643
0.000000000000 2.386766012830 1.729854825170 | 3.681777820765
2.067000000000 1.193383006415 4.922145174830 | 3.681777820765
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 26.608000000000 | 23.001046659363 <- sum
maximal spread = 3.681777820765
####################################################################################################
####################################################################################################
Iteration 0 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 0.000000000000 | 0.478929974197
0.000000000000 0.000000000000 0.000000000000 | 0.469563282099
0.000000000000 0.000000000000 0.000000000000 | 0.469563282099
0.000000000000 0.000000000000 0.000000000000 | 0.454607455891
0.000000000000 0.000000000000 0.000000000000 | 0.454607455891
0.715265049983 2.799724481980 1.679302428793 | 2.205936575534
-0.715265049983 2.799724481980 1.679302428793 | 2.205936575534
-0.000000000000 1.560849074530 1.679302428793 | 2.205936575534
1.351734950017 0.780424537265 4.972697571207 | 2.205936575534
2.782265049983 0.780424537265 4.972697571207 | 2.205936575534
2.067000000000 2.019299944714 4.972697571207 | 2.205936575534
0.000000000000 2.386766012830 1.730315184839 | 3.569118724074
2.067000000000 1.193383006415 4.921684815161 | 3.569118724074
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 26.608000000000 | 22.701128351528 <- sum
maximal spread = 3.569118724074
standard deviation = 0.0
####################################################################################################
####################################################################################################
Iteration 50 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 0.000000000000 | 0.453286739562
0.000000000000 0.000000000000 0.000000000000 | 0.473931288465
0.000000000000 0.000000000000 0.000000000000 | 0.473931288465
0.000000000000 0.000000000000 0.000000000000 | 0.451955985201
0.000000000000 0.000000000000 0.000000000000 | 0.451955985201
0.711471235888 2.797534122391 1.688699327935 | 2.082541481473
-0.711471235888 2.797534122391 1.688699327935 | 2.082541481473
0.000000000000 1.565229793708 1.688699327935 | 2.082541481473
1.355528764112 0.782614896854 4.963300672065 | 2.082541481473
2.778471235888 0.782614896854 4.963300672065 | 2.082541481473
2.067000000000 2.014919225537 4.963300672065 | 2.082541481473
0.000000000000 2.386766012830 1.747860130855 | 3.450336620111
2.067000000000 1.193383006415 4.904139869145 | 3.450336620111
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 26.608000000000 | 21.700983415953 <- sum
maximal spread = 3.450336620111
standard deviation = 0.00032356493497899715
####################################################################################################
####################################################################################################
Iteration 100 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 0.000000000000 | 0.453820475336
0.000000000000 0.000000000000 0.000000000000 | 0.479557103935
0.000000000000 0.000000000000 0.000000000000 | 0.479557103935
0.000000000000 0.000000000000 0.000000000000 | 0.453683180782
0.000000000000 0.000000000000 0.000000000000 | 0.453683180782
0.711200009612 2.797377529827 1.680969007833 | 2.082236906048
-0.711200009612 2.797377529827 1.680969007833 | 2.082236906048
-0.000000000000 1.565542978836 1.680969007833 | 2.082236906048
1.355799990388 0.782771489418 4.971030992167 | 2.082236906048
2.778200009612 0.782771489418 4.971030992167 | 2.082236906048
2.067000000000 2.014606040409 4.971030992167 | 2.082236906048
-0.000000000000 2.386766012830 1.772146727899 | 3.442205183445
2.067000000000 1.193383006415 4.879853272101 | 3.442205183445
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 26.608000000000 | 21.698132847948 <- sum
maximal spread = 3.442205183445
standard deviation = 0.0004722647203498406
####################################################################################################
####################################################################################################
Iteration 150 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 0.000000000000 | 0.453633282234
0.000000000000 0.000000000000 0.000000000000 | 0.480067018952
0.000000000000 0.000000000000 0.000000000000 | 0.480067018952
0.000000000000 0.000000000000 0.000000000000 | 0.453623536637
0.000000000000 0.000000000000 0.000000000000 | 0.453623536637
0.710475947474 2.796959492357 1.669670390075 | 2.085695692500
-0.710475947474 2.796959492357 1.669670390075 | 2.085695692500
0.000000000000 1.566379053776 1.669670390075 | 2.085695692500
1.356524052526 0.783189526888 4.982329609925 | 2.085695692500
2.777475947474 0.783189526888 4.982329609925 | 2.085695692500
2.067000000000 2.013769965469 4.982329609925 | 2.085695692500
-0.000000000000 2.386766012830 1.807330449040 | 3.428604694868
2.067000000000 1.193383006415 4.844669550960 | 3.428604694868
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 26.608000000000 | 21.692397938146 <- sum
maximal spread = 3.428604694868
standard deviation = 0.0006788117298202646
####################################################################################################
####################################################################################################
Iteration 200 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 0.000000000000 | 0.453334337312
0.000000000000 0.000000000000 0.000000000000 | 0.480473130458
0.000000000000 0.000000000000 0.000000000000 | 0.480473130458
0.000000000000 0.000000000000 0.000000000000 | 0.453464224943
0.000000000000 0.000000000000 0.000000000000 | 0.453464224943
0.708889448927 2.796043526994 1.653672955618 | 2.092459498606
-0.708889448927 2.796043526994 1.653672955618 | 2.092459498606
-0.000000000000 1.568210984503 1.653672955618 | 2.092459498606
1.358110551073 0.784105492251 4.998327044382 | 2.092459498606
2.775889448927 0.784105492251 4.998327044382 | 2.092459498606
2.067000000000 2.011938034742 4.998327044382 | 2.092459498606
-0.000000000000 2.386766012830 1.857051355735 | 3.402228011503
2.067000000000 1.193383006415 4.794948644265 | 3.402228011503
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 26.608000000000 | 21.680422062756 <- sum
maximal spread = 3.402228011503
standard deviation = 0.0009424543163257928
####################################################################################################
####################################################################################################
Iteration 250 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 0.000000000000 | 0.452940219315
0.000000000000 0.000000000000 0.000000000000 | 0.480976129644
0.000000000000 0.000000000000 0.000000000000 | 0.480976129644
0.000000000000 0.000000000000 0.000000000000 | 0.453286401798
0.000000000000 0.000000000000 0.000000000000 | 0.453286401798
0.705669179877 2.794184303791 1.632126188959 | 2.104977550522
-0.705669179877 2.794184303791 1.632126188959 | 2.104977550522
-0.000000000000 1.571929430908 1.632126188959 | 2.104977550522
1.361330820123 0.785964715454 5.019873811041 | 2.104977550522
2.772669179877 0.785964715454 5.019873811041 | 2.104977550522
2.067000000000 2.008219588336 5.019873811041 | 2.104977550522
0.000000000000 2.386766012830 1.923817087720 | 3.353167399203
2.067000000000 1.193383006415 4.728182912280 | 3.353167399203
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 26.608000000000 | 21.657665383739 <- sum
maximal spread = 3.353167399203
standard deviation = 0.0012227663963830747
####################################################################################################
####################################################################################################
Final state (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 0.000000000000 | 0.452480642054
0.000000000000 0.000000000000 0.000000000000 | 0.481524374305
0.000000000000 0.000000000000 0.000000000000 | 0.481524374305
0.000000000000 0.000000000000 0.000000000000 | 0.453141933770
0.000000000000 0.000000000000 0.000000000000 | 0.453141933770
0.699985700042 2.790902945178 1.606193018682 | 2.125716972317
-0.699985700042 2.790902945178 1.606193018682 | 2.125716972317
0.000000000000 1.578492148134 1.606193018682 | 2.125716972317
1.367014299958 0.789246074067 5.045806981318 | 2.125716972317
2.766985700042 0.789246074067 5.045806981318 | 2.125716972317
2.067000000000 2.001656871111 5.045806981318 | 2.125716972317
0.000000000000 2.386766012830 2.003757873952 | 3.272761655937
2.067000000000 1.193383006415 4.648242126048 | 3.272761655937
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 26.608000000000 | 21.621638403977 <- sum
maximal spread = 3.272761655937
standard deviation = 0.0016237004013601678
####################################################################################################
time for creating wannierizer 0.92922043800354
time for iterations 12.690051317214966
time for updating 9.780243873596191
total time for wannierization 15.362960815429688
saving to wannier_soc-spin-0.chk :
Warning: symmetrization did not converge in 10 iterations, final changes 0.003065492738868214, 2.2868100709761296e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.004130995006472277, 5.968028382946875e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=1, kpt=1, [0. 0. 0.25] with 12 symmetries, max error in included blocks: 1.2019057696113271e-06 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 37 [55:56] is 1.2019057696113271e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
Warning: symmetrization did not converge in 10 iterations, final changes 5.7644955675923095e-05, 1.2888733563044084e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 3.246611072795757e-05, 1.1707708926743101e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.0002399293011803818, 4.295625719062655e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.0001409118510938896, 2.078658940906133e-05; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.0019627241924937253, 0.0006063250915731706; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=8, kpt=8, [0. 0.33333333 0.25 ] with 4 symmetries, max error in included blocks: 1.5643795696135995e-06 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 49 [54:55] is 1.5643795696135995e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 50 [54:55] is 9.259808422651666e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
Warning: symmetrization did not converge in 10 iterations, final changes 0.0008372957054689798, 0.0002257377502176513; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.0008332065611458957, 0.00020108495848781413; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=10, kpt=10, [0. 0.33333333 0.75 ] with 4 symmetries, max error in included blocks: 9.259808422651666e-06 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
Warning: symmetrization did not converge in 10 iterations, final changes 0.00037997766440883864, 0.00011443188752756144; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Warning: symmetrization did not converge in 10 iterations, final changes 0.0009221888765184094, 0.00023263480129269995; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=13, kpt=13, [0. 0.5 0.5] with 8 symmetries, max error in included blocks: 7.865475366342547e-06 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 50 [54:55] is 1.9172723939621294e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 51 [55:56] is 7.865475366342547e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
Warning: symmetrization did not converge in 10 iterations, final changes 0.003595245080980007, 0.00016103523832065652; probably the input data are not perfectly symmetrizable, or the provided projections are notcompatible with the irreps of the DFT bands.
Symmetrizer_Uirr initialized for ikirr=21, kpt=21, [0.33333333 0.33333333 0. ] with 12 symmetries, max error in included blocks: 0.002382115755663211 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
Symmetrizer_Uirr initialized for ikirr=22, kpt=22, [0.33333333 0.33333333 0.25 ] with 6 symmetries, max error in included blocks: 1.3941251176678493e-05 ; excluded bands are [56 57 58 59] out of 60 total bands (accuracy threshold 1e-06)
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 13 [22:24] is 1.0023166867707734e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 32 [51:53] is 1.6929346154941593e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 33 [53:54] is 0.0002481579806393198, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 34 [54:56] is 0.002382115755663211, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 15 [25:27] is 1.3941251176678493e-05, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 33 [53:55] is 1.1602455039699551e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
/home/stepan/github/wannier-berri/wannierberri/symmetry/sawf_kirr.py:163: UserWarning: Warning: max error in block 34 [55:56] is 1.4802404153727076e-06, exceeding threshold 1e-06, and this is not among the upper bands(56:60) bands, this may indicate inaccuracy in the input data
warnings.warn(f"Warning: max error in block {i} [{start}:{end}] is {max_error_in_blocks[i]}, exceeding threshold {accuracy_threshold}, and this is not among the upper"
####################################################################################################
starting WFs
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 3.326000000000 | 0.484461313215
0.000000000000 0.000000000000 3.326000000000 | 0.473499120352
0.000000000000 0.000000000000 3.326000000000 | 0.473499120352
0.000000000000 0.000000000000 3.326000000000 | 0.457942503894
0.000000000000 0.000000000000 3.326000000000 | 0.457942503894
0.715030052524 2.799588806134 1.647908921279 | 2.215017239905
-0.715030052524 2.799588806134 1.647908921279 | 2.215017239905
-0.000000000000 1.561120426223 1.647908921279 | 2.215017239905
1.351969947476 0.780560213111 5.004091078721 | 2.215017239905
2.782030052524 0.780560213111 5.004091078721 | 2.215017239905
2.067000000000 2.019028593022 5.004091078721 | 2.215017239905
0.000000000000 2.386766012830 1.596164169636 | 3.681800051957
2.067000000000 1.193383006415 5.055835830364 | 3.681800051957
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 43.238000000000 | 23.001048105048 <- sum
maximal spread = 3.681800051957
####################################################################################################
####################################################################################################
Iteration 0 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 3.326000000000 | 0.478901719527
0.000000000000 0.000000000000 3.326000000000 | 0.469554312720
0.000000000000 0.000000000000 3.326000000000 | 0.469554312720
0.000000000000 0.000000000000 3.326000000000 | 0.454610950562
0.000000000000 0.000000000000 3.326000000000 | 0.454610950562
0.715264959855 2.799724429944 1.646649055395 | 2.205935654503
-0.715264959855 2.799724429944 1.646649055395 | 2.205935654503
0.000000000000 1.560849178601 1.646649055395 | 2.205935654503
1.351735040145 0.780424589301 5.005350944605 | 2.205935654503
2.782264959855 0.780424589301 5.005350944605 | 2.205935654503
2.067000000000 2.019299840644 5.005350944605 | 2.205935654503
0.000000000000 2.386766012830 1.595704837124 | 3.569143177662
2.067000000000 1.193383006415 5.056295162876 | 3.569143177662
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 43.238000000000 | 22.701132528431 <- sum
maximal spread = 3.569143177662
standard deviation = 0.0
####################################################################################################
####################################################################################################
Iteration 50 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 3.326000000000 | 0.453297909835
0.000000000000 0.000000000000 3.326000000000 | 0.473936629272
0.000000000000 0.000000000000 3.326000000000 | 0.473936629272
0.000000000000 0.000000000000 3.326000000000 | 0.451967379109
0.000000000000 0.000000000000 3.326000000000 | 0.451967379109
0.711471146797 2.797534070954 1.637264963018 | 2.082552662484
-0.711471146797 2.797534070954 1.637264963018 | 2.082552662484
0.000000000000 1.565229896583 1.637264963018 | 2.082552662484
1.355528853203 0.782614948291 5.014735036982 | 2.082552662484
2.778471146797 0.782614948291 5.014735036982 | 2.082552662484
2.067000000000 2.014919122662 5.014735036982 | 2.082552662484
-0.000000000000 2.386766012830 1.578229395843 | 3.450333803712
2.067000000000 1.193383006415 5.073770604157 | 3.450333803712
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 43.238000000000 | 21.701089508925 <- sum
maximal spread = 3.450333803712
standard deviation = 0.0003223108567665236
####################################################################################################
####################################################################################################
Iteration 100 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 3.326000000000 | 0.453833291086
0.000000000000 0.000000000000 3.326000000000 | 0.479562753095
0.000000000000 0.000000000000 3.326000000000 | 0.479562753095
0.000000000000 0.000000000000 3.326000000000 | 0.453695094719
0.000000000000 0.000000000000 3.326000000000 | 0.453695094719
0.711201943276 2.797378646228 1.644965422100 | 2.082239390273
-0.711201943276 2.797378646228 1.644965422100 | 2.082239390273
-0.000000000000 1.565540746033 1.644965422100 | 2.082239390273
1.355798056724 0.782770373017 5.007034577900 | 2.082239390273
2.778201943276 0.782770373017 5.007034577900 | 2.082239390273
2.067000000000 2.014608273212 5.007034577900 | 2.082239390273
0.000000000000 2.386766012830 1.554036021193 | 3.442238460053
2.067000000000 1.193383006415 5.097963978807 | 3.442238460053
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 43.238000000000 | 21.698262248459 <- sum
maximal spread = 3.442238460053
standard deviation = 0.0004704692868973237
####################################################################################################
####################################################################################################
Iteration 150 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 3.326000000000 | 0.453646685793
0.000000000000 0.000000000000 3.326000000000 | 0.480072037678
0.000000000000 0.000000000000 3.326000000000 | 0.480072037678
0.000000000000 0.000000000000 3.326000000000 | 0.453635545847
0.000000000000 0.000000000000 3.326000000000 | 0.453635545847
0.710482619778 2.796963344613 1.656221931878 | 2.085679111477
-0.710482619778 2.796963344613 1.656221931878 | 2.085679111477
-0.000000000000 1.566371349263 1.656221931878 | 2.085679111477
1.356517380222 0.783185674632 4.995778068122 | 2.085679111477
2.777482619778 0.783185674632 4.995778068122 | 2.085679111477
2.067000000000 2.013777669982 4.995778068122 | 2.085679111477
-0.000000000000 2.386766012830 1.518983051205 | 3.428715850818
2.067000000000 1.193383006415 5.133016948795 | 3.428715850818
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 43.238000000000 | 21.692568223343 <- sum
maximal spread = 3.428715850818
standard deviation = 0.0006763520025546952
####################################################################################################
####################################################################################################
Iteration 200 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 3.326000000000 | 0.453348363669
0.000000000000 0.000000000000 3.326000000000 | 0.480477363364
0.000000000000 0.000000000000 3.326000000000 | 0.480477363364
0.000000000000 0.000000000000 3.326000000000 | 0.453476267305
0.000000000000 0.000000000000 3.326000000000 | 0.453476267305
0.708906575689 2.796053415134 1.672164175361 | 2.092404568225
-0.708906575689 2.796053415134 1.672164175361 | 2.092404568225
0.000000000000 1.568191208222 1.672164175361 | 2.092404568225
1.358093424311 0.784095604111 4.979835824639 | 2.092404568225
2.775906575689 0.784095604111 4.979835824639 | 2.092404568225
2.067000000000 2.011957811023 4.979835824639 | 2.092404568225
-0.000000000000 2.386766012830 1.469432736821 | 3.402495069166
2.067000000000 1.193383006415 5.182567263179 | 3.402495069166
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 43.238000000000 | 21.680673172691 <- sum
maximal spread = 3.402495069166
standard deviation = 0.0009394135574282215
####################################################################################################
####################################################################################################
Iteration 250 (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 3.326000000000 | 0.452954771990
0.000000000000 0.000000000000 3.326000000000 | 0.480979737692
0.000000000000 0.000000000000 3.326000000000 | 0.480979737692
0.000000000000 0.000000000000 3.326000000000 | 0.453298195175
0.000000000000 0.000000000000 3.326000000000 | 0.453298195175
0.705706231561 2.794205695591 1.693649067508 | 2.104853949448
-0.705706231561 2.794205695591 1.693649067508 | 2.104853949448
-0.000000000000 1.571886647308 1.693649067508 | 2.104853949448
1.361293768439 0.785943323654 4.958350932492 | 2.104853949448
2.772706231561 0.785943323654 4.958350932492 | 2.104853949448
2.067000000000 2.008262371936 4.958350932492 | 2.104853949448
0.000000000000 2.386766012830 1.402856340243 | 3.353707842714
2.067000000000 1.193383006415 5.249143659757 | 3.353707842714
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 43.238000000000 | 21.658050019840 <- sum
maximal spread = 3.353707842714
standard deviation = 0.0012197987174394841
####################################################################################################
####################################################################################################
Final state (from wannierizer)
----------------------------------------------------------------------------------------------------
wannier centers and spreads
----------------------------------------------------------------------------------------------------
0.000000000000 0.000000000000 3.326000000000 | 0.452495252047
0.000000000000 0.000000000000 3.326000000000 | 0.481528105841
0.000000000000 0.000000000000 3.326000000000 | 0.481528105841
0.000000000000 0.000000000000 3.326000000000 | 0.453152907533
0.000000000000 0.000000000000 3.326000000000 | 0.453152907533
0.700052056937 2.790941256349 1.719535024031 | 2.125497055898
-0.700052056937 2.790941256349 1.719535024031 | 2.125497055898
0.000000000000 1.578415525791 1.719535024031 | 2.125497055898
1.366947943063 0.789207762896 4.932464975969 | 2.125497055898
2.767052056937 0.789207762896 4.932464975969 | 2.125497055898
2.067000000000 2.001733493454 4.932464975969 | 2.125497055898
0.000000000000 2.386766012830 1.323056151525 | 3.273666498815
2.067000000000 1.193383006415 5.328943848475 | 3.273666498815
----------------------------------------------------------------------------------------------------
8.268000000000 14.320596076979 43.238000000000 | 21.622172611814 <- sum
maximal spread = 3.273666498815
standard deviation = 0.001617465319686861
####################################################################################################
time for creating wannierizer 1.2086689472198486
time for iterations 13.381288766860962
time for updating 10.23925495147705
total time for wannierization 16.50211238861084
saving to wannier_soc-spin-1.chk :
Create the System object to be used in WannierBerri calculations
It is called “System_w90” class, although it now does not use the w90 wannierization, but the class is the same.
Unlike the non-SOC systems, here we also specify the direction of the magnetization. We can later easily chanbge this direction without redoing the wannierization.
[12]:
from wannierberri.system.system_soc import SystemSOC
theta = 90
phi = 90
system_soc = SystemSOC.from_wannierdata(wandata=wandata, berry=True, silent=False)
system_soc.set_soc_axis(theta=theta, phi=phi, alpha_soc=1.0, units="degrees")
system_soc.save_npz("system_soc")
irreducible : True, symmetrize set to True
setting Rvec
setting AA..
setting AA - OK
Real-space lattice:
[[ 4.134 0. 0. ]
[-2.067 3.58014902 0. ]
[ 0. 0. 6.652 ]]
Number of wannier functions: 13
Number of R points: 267
Recommended size of FFT grid [6 6 4]
num_blocks_left = 3, num_blocks_right = 3
number o R-vectors before symmetrization: 267
number o R-vectors after symmetrization: 573
irreducible : True, symmetrize set to True
setting Rvec
setting AA..
setting AA - OK
Real-space lattice:
[[ 4.134 0. 0. ]
[-2.067 3.58014902 0. ]
[ 0. 0. 6.652 ]]
Number of wannier functions: 13
Number of R points: 267
Recommended size of FFT grid [6 6 4]
num_blocks_left = 3, num_blocks_right = 3
number o R-vectors before symmetrization: 267
number o R-vectors after symmetrization: 425
nspin in SOC: 2
setting Rvec
/home/stepan/github/wannier-berri/wannierberri/utility.py:57: UserWarning: usually need to provide either with real or reciprocal lattice.If you only want to generate a random symmetric tensor - that it fine
warnings.warn("usually need to provide either with real or reciprocal lattice."
using magmoms
[[ 0.00000000e+00 0.00000000e+00 4.40436811e+00]
[-0.00000000e+00 -0.00000000e+00 -4.40436811e+00]
[-0.00000000e+00 -0.00000000e+00 -2.92436898e-10]
[-0.00000000e+00 -0.00000000e+00 -2.92436898e-10]]
num_blocks_left = 3, num_blocks_right = 3
num_blocks_left = 3, num_blocks_right = 3
num_blocks_left = 3, num_blocks_right = 3
using magmoms
[[ 0.00000000e+00 0.00000000e+00 4.40436811e+00]
[-0.00000000e+00 -0.00000000e+00 -4.40436811e+00]
[-0.00000000e+00 -0.00000000e+00 -2.92436898e-10]
[-0.00000000e+00 -0.00000000e+00 -2.92436898e-10]]
using magmoms
[[ 2.69689765e-16 4.40436811e+00 2.69689765e-16]
[-2.69689765e-16 -4.40436811e+00 -2.69689765e-16]
[-1.79065955e-26 -2.92436898e-10 -1.79065955e-26]
[-1.79065955e-26 -2.92436898e-10 -1.79065955e-26]]
Saving SystemSOC to system_soc
saving system of class SystemSOC to system_soc
properties: ['num_wann', 'real_lattice', 'iRvec', 'periodic', 'is_phonon', 'wannier_centers_cart', 'pointgroup', 'cell']
saving num_wann to system_soc/num_wann.npz
saving real_lattice to system_soc/real_lattice.npz
saving iRvec to system_soc/iRvec.npz
saving periodic to system_soc/periodic.npz
saving is_phonon to system_soc/is_phonon.npz
saving wannier_centers_cart to system_soc/wannier_centers_cart.npz
saving pointgroup to system_soc/pointgroup.npz
saving cell to system_soc/cell.npz
saving system of class System_R to system_soc/system_up
properties: ['num_wann', 'real_lattice', 'iRvec', 'periodic', 'is_phonon', 'wannier_centers_cart', 'pointgroup']
saving num_wann
saving num_wann to system_soc/system_up/num_wann.npz
- Ok!
saving real_lattice
saving real_lattice to system_soc/system_up/real_lattice.npz
- Ok!
saving iRvec
saving iRvec to system_soc/system_up/iRvec.npz
- Ok!
saving periodic
saving periodic to system_soc/system_up/periodic.npz
- Ok!
saving is_phonon
saving is_phonon to system_soc/system_up/is_phonon.npz
- Ok!
saving wannier_centers_cart
saving wannier_centers_cart to system_soc/system_up/wannier_centers_cart.npz
- Ok!
saving pointgroup
saving pointgroup to system_soc/system_up/pointgroup.npz
- Ok!
saving Ham - Ok!
saving AA - Ok!
saving system of class System_R to system_soc/system_down
properties: ['num_wann', 'real_lattice', 'iRvec', 'periodic', 'is_phonon', 'wannier_centers_cart', 'pointgroup']
saving num_wann
saving num_wann to system_soc/system_down/num_wann.npz
- Ok!
saving real_lattice
saving real_lattice to system_soc/system_down/real_lattice.npz
- Ok!
saving iRvec
saving iRvec to system_soc/system_down/iRvec.npz
- Ok!
saving periodic
saving periodic to system_soc/system_down/periodic.npz
- Ok!
saving is_phonon
saving is_phonon to system_soc/system_down/is_phonon.npz
- Ok!
saving wannier_centers_cart
saving wannier_centers_cart to system_soc/system_down/wannier_centers_cart.npz
- Ok!
saving pointgroup
saving pointgroup to system_soc/system_down/pointgroup.npz
- Ok!
saving Ham - Ok!
saving AA - Ok!
Compute the wannierized bandstructure with SOC along the high-symmetry path
[15]:
from wannierberri.grid import Path
from wannierberri.evaluate_k import evaluate_k_path
from wannierberri.system.system_soc import SystemSOC
system_soc = SystemSOC.from_npz("system_soc")
kz = 0.35 / (2*np.pi/c)
path = Path.from_nodes(real_lattice=lattice,
nodes=[
[2 / 3, -1 / 3, 0],
[0, 0, 0],
[-2 / 3, 1 / 3, 0],
None,
[-0.5, 0, kz],
[0, 0, kz],
[0.5, 0, kz],
],
labels=[r"${\rm K}\leftarrow$",
r"$\Gamma$",
r"$\rightarrow{\rm K}$",
r"$\overline{\rm M}\leftarrow$",
r"$\overline{\Gamma}$",
r"$\rightarrow\overline{\rm M}$"],
length=300) # length [ Ang] ~= 2*pi/dk
bands_wannier_soc= evaluate_k_path(system_soc, path=path, quantities=["spin"])
bands_wannier_up = evaluate_k_path(system_soc.system_up, path=path)
bands_wannier_dw = evaluate_k_path(system_soc.system_down, path=path)
loading real_lattice - Ok!
loading wannier_centers_cart - Ok!
loading pointgroup - Ok!
loading iRvec - Ok!
loading periodic - Ok!
loading is_phonon - Ok!
loading atom_labels - Ok!
loading num_wann - Ok!
loading positions - Ok!
loading R_matrix AA - Ok!
loading R_matrix Ham - Ok!
loading real_lattice - Ok!
loading wannier_centers_cart - Ok!
loading pointgroup - Ok!
loading iRvec - Ok!
loading periodic - Ok!
loading is_phonon - Ok!
loading atom_labels - Ok!
loading num_wann - Ok!
loading positions - Ok!
loading R_matrix AA - Ok!
loading R_matrix Ham - Ok!
Starting run()
Using the follwing calculators :
############################################################
'tabulate' : <wannierberri.calculators.tabulate.TabulatorAll object at 0x714803587530> :
TabulatorAll - a pack of all k-resolved calculators (Tabulators)
Includes the following tabulators :
--------------------------------------------------
"spin" : <wannierberri.calculators.tabulate.Spin object at 0x714801f2a960> : Spin expectation :math:` \langle u | \mathbf{\sigma} | u \rangle`
"Energy" : <wannierberri.calculators.tabulate.Energy object at 0x7148009e5a30> : calculator not described
--------------------------------------------------
############################################################
Calculation along a path - checking calculators for compatibility
tabulate <wannierberri.calculators.tabulate.TabulatorAll object at 0x714803587530>
All calculators are compatible
Symmetrization switched off for Path
Grid is regular
The set of k points is a Path() with 182 points and labels {0: '${\\rm K}\\leftarrow$', 48: '$\\Gamma$', 96: '$\\rightarrow{\\rm K}$', 97: '$\\overline{\\rm M}\\leftarrow$', 139: '$\\overline{\\Gamma}$', 181: '$\\rightarrow\\overline{\\rm M}$'}
generating K_list
Done
Done, sum of weights:182.0
############################################################
Iteration 0 out of 0
processing 182 K points : using 8 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
/home/stepan/github/wannier-berri/wannierberri/utility.py:57: UserWarning: usually need to provide either with real or reciprocal lattice.If you only want to generate a random symmetric tensor - that it fine
warnings.warn("usually need to provide either with real or reciprocal lattice."
/home/stepan/github/wannier-berri/wannierberri/grid/path.py:272: UserWarning: symmetry is not used for a tabulation along path
warnings.warn("symmetry is not used for a tabulation along path")
time for processing 182 K-points on 8 processes: 1.3089 ; per K-point 0.0072 ; proc-sec per K-point 0.0575
time1 = 4.76837158203125e-07
Totally processed 182 K-points
run() finished
Starting run()
Using the follwing calculators :
############################################################
'tabulate' : <wannierberri.calculators.tabulate.TabulatorAll object at 0x714803587530> :
TabulatorAll - a pack of all k-resolved calculators (Tabulators)
Includes the following tabulators :
--------------------------------------------------
"Energy" : <wannierberri.calculators.tabulate.Energy object at 0x7148009db440> : calculator not described
--------------------------------------------------
############################################################
Calculation along a path - checking calculators for compatibility
tabulate <wannierberri.calculators.tabulate.TabulatorAll object at 0x714803587530>
All calculators are compatible
Symmetrization switched off for Path
Grid is regular
The set of k points is a Path() with 182 points and labels {0: '${\\rm K}\\leftarrow$', 48: '$\\Gamma$', 96: '$\\rightarrow{\\rm K}$', 97: '$\\overline{\\rm M}\\leftarrow$', 139: '$\\overline{\\Gamma}$', 181: '$\\rightarrow\\overline{\\rm M}$'}
generating K_list
Done
Done, sum of weights:182.0
############################################################
Iteration 0 out of 0
processing 182 K points : using 8 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 182 K-points on 8 processes: 0.1635 ; per K-point 0.0009 ; proc-sec per K-point 0.0072
time1 = 2.384185791015625e-07
Totally processed 182 K-points
run() finished
Starting run()
Using the follwing calculators :
############################################################
'tabulate' : <wannierberri.calculators.tabulate.TabulatorAll object at 0x714803587530> :
TabulatorAll - a pack of all k-resolved calculators (Tabulators)
Includes the following tabulators :
--------------------------------------------------
"Energy" : <wannierberri.calculators.tabulate.Energy object at 0x714802270590> : calculator not described
--------------------------------------------------
############################################################
Calculation along a path - checking calculators for compatibility
tabulate <wannierberri.calculators.tabulate.TabulatorAll object at 0x714803587530>
All calculators are compatible
Symmetrization switched off for Path
Grid is regular
The set of k points is a Path() with 182 points and labels {0: '${\\rm K}\\leftarrow$', 48: '$\\Gamma$', 96: '$\\rightarrow{\\rm K}$', 97: '$\\overline{\\rm M}\\leftarrow$', 139: '$\\overline{\\Gamma}$', 181: '$\\rightarrow\\overline{\\rm M}$'}
generating K_list
Done
Done, sum of weights:182.0
############################################################
Iteration 0 out of 0
processing 182 K points : using 8 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 182 K-points on 8 processes: 0.1397 ; per K-point 0.0008 ; proc-sec per K-point 0.0061
time1 = 0.0
Totally processed 182 K-points
run() finished
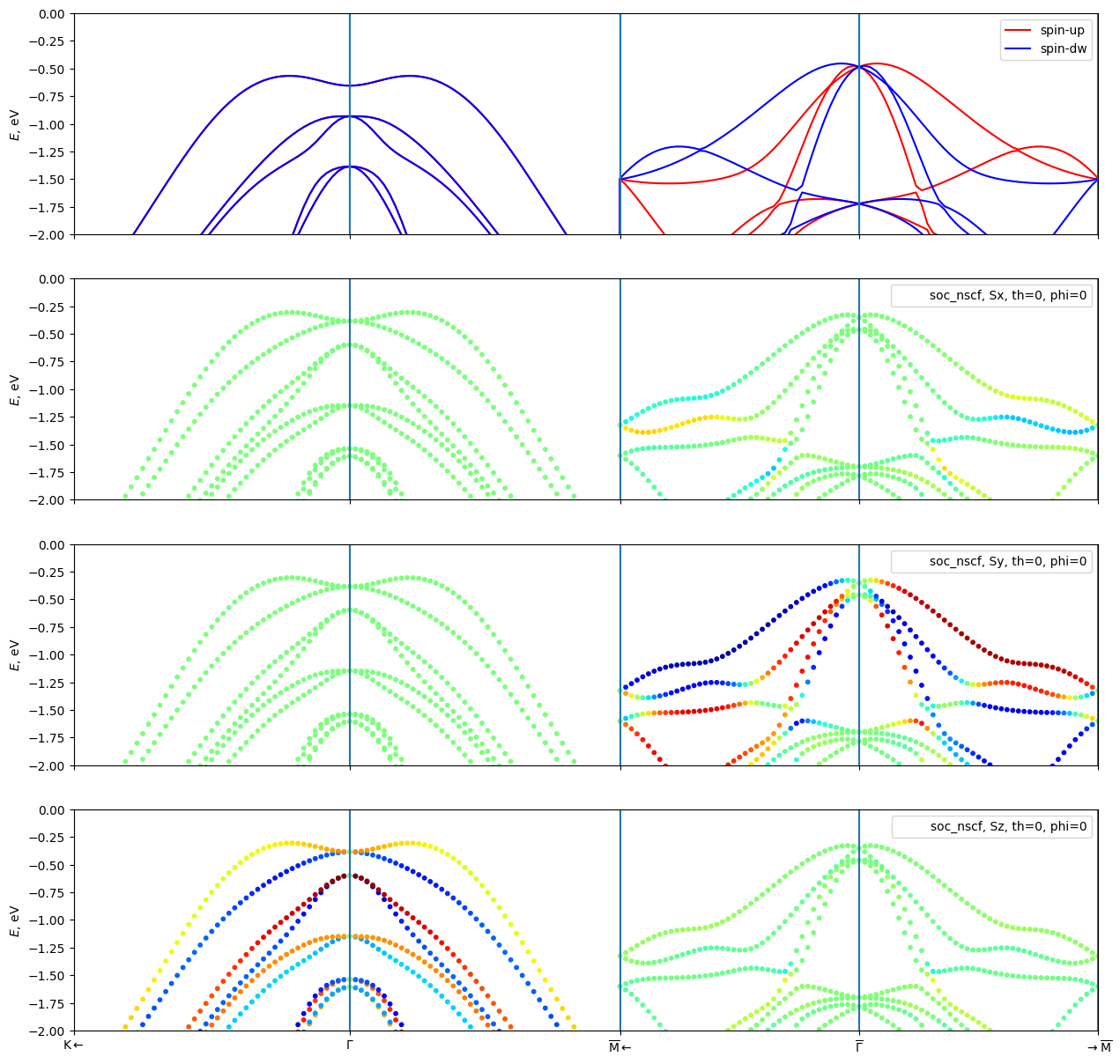
Plot the Bandstructure with and without SOC
[16]:
from matplotlib import pyplot as plt
EF = 6.885145845031927
fig, axes = plt.subplots(4, 1, sharey=True, sharex=True, figsize=(15, 15))
for i in range(3):
component = "xyz"[i]
bands_wannier_soc.plot_path_fat(path=path,
Eshift=EF,
quantity="spin",
component=component,
mode="color",
label=f"soc_nscf, S{component}, th={theta}, phi={phi}",
axes=axes[1 + i],
fatmax=10,
linecolor="orange",
close_fig=False,
show_fig=False,
kwargs_line=dict(linestyle='-', lw=0.0),
)
bands_wannier_up.plot_path_fat(path=path,
Eshift=EF,
axes=axes[0],
label="spin-up",
linecolor="red",
close_fig=False,
show_fig=False,
)
bands_wannier_dw.plot_path_fat(path=path,
label="spin-dw",
axes=axes[0],
Eshift=EF,
linecolor="blue",
close_fig=False,
show_fig=False,)
plt.ylim(-2,0)
plt.show()
AHC
magnetic moments along “y” direction
Below we evaluate only the “internal” (which depend only on the Hamiltonian) contributions to the anomalous Hall conductivity (AHC), how to include the “external” contributions is explained in the previous tutorial on bcc Fe.
[2]:
import os
import numpy as np
import wannierberri as wb
from wannierberri.system.system_soc import SystemSOC
system_soc_y = SystemSOC.from_npz("system_soc")
theta=90
phi=90
system_soc_y.set_soc_axis(theta=theta, phi=phi, units="degrees")
grid = wb.grid.Grid(system_soc_y, NK=200)
EF = 6.885145845031927
tetra = False
Efermi = np.linspace(EF - 2, EF, 1001)
os.makedirs("results", exist_ok=True)
result_int = wb.run(system_soc_y,
grid=grid,
fout_name=f"results/{seed}-soc-y",
calculators={"ahc_int": wb.calculators.static.AHC(Efermi=Efermi, tetra=tetra, kwargs_formula={"external_terms": False})},
adpt_num_iter=20,
)
loading real_lattice - Ok!
loading wannier_centers_cart - Ok!
loading pointgroup - Ok!
loading iRvec - Ok!
loading periodic - Ok!
loading is_phonon - Ok!
loading atom_labels - Ok!
loading num_wann - Ok!
loading positions - Ok!
loading R_matrix AA - Ok!
loading R_matrix Ham - Ok!
loading real_lattice - Ok!
loading wannier_centers_cart - Ok!
loading pointgroup - Ok!
loading iRvec - Ok!
loading periodic - Ok!
loading is_phonon - Ok!
loading atom_labels - Ok!
loading num_wann - Ok!
loading positions - Ok!
loading R_matrix AA - Ok!
loading R_matrix Ham - Ok!
/home/stepan/github/wannier-berri/wannierberri/utility.py:57: UserWarning: usually need to provide either with real or reciprocal lattice.If you only want to generate a random symmetric tensor - that it fine
warnings.warn("usually need to provide either with real or reciprocal lattice."
using magmoms
[[ 2.69689765e-16 4.40436811e+00 2.69689765e-16]
[-2.69689765e-16 -4.40436811e+00 -2.69689765e-16]
[-1.79065955e-26 -2.92436898e-10 -1.79065955e-26]
[-1.79065955e-26 -2.92436898e-10 -1.79065955e-26]]
Minimal symmetric FFT grid : [11 11 7]
Starting run()
Using the follwing calculators :
############################################################
'ahc_int' : <wannierberri.calculators.static.AHC object at 0x75643fdae240> : Anomalous Hall conductivity (:math:`s^3 \cdot A^2 / (kg \cdot m^3) = S/m`)
| With Fermi sea integral Eq(11) in `Ref <https://www.nature.com/articles/s41524-021-00498-5>`__
| Output: :math:`O = -e^2/\hbar \int [dk] \Omega f`
| Instruction: :math:`j_\alpha = \sigma_{\alpha\beta} E_\beta = \epsilon_{\alpha\beta\delta} O_\delta E_\beta`
############################################################
Calculation on grid - checking calculators for compatibility
ahc_int <wannierberri.calculators.static.AHC object at 0x75643fdae240>
All calculators are compatible
Grid is regular
The set of k points is a Grid() with NKdiv=[10 10 25], NKFFT=[20 20 8], NKtot=[200 200 200]
generating K_list
Done in 0.006664276123046875 s
excluding symmetry-equivalent K-points from initial grid
Done in 0.05006265640258789 s
K_list contains 403 Irreducible points(16.12%) out of initial 10x10x25=2500 grid
Done, sum of weights:0.999999999999999
############################################################
Iteration 0 out of 20
processing 403 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
16 11.1 267.9 279.0
32 17.3 200.4 217.7
48 24.3 179.9 204.2
64 31.0 164.3 195.3
80 37.8 152.8 190.6
96 44.4 142.0 186.4
112 50.9 132.2 183.1
128 57.2 122.8 180.0
144 63.6 114.5 178.1
160 71.0 107.9 178.9
176 78.1 100.7 178.9
192 84.6 93.0 177.7
208 90.9 85.2 176.1
224 98.1 78.4 176.4
240 105.4 71.6 177.0
256 112.9 64.8 177.8
272 119.9 57.7 177.6
288 126.9 50.7 177.5
304 132.9 43.3 176.2
320 140.8 36.5 177.3
336 147.2 29.3 176.5
352 155.8 22.6 178.4
368 161.7 15.4 177.1
384 167.7 8.3 176.0
400 174.5 1.3 175.8
time for processing 403 K-points on 16 processes: 175.9319 ; per K-point 0.4366 ; proc-sec per K-point 6.9849
time1 = 4.76837158203125e-07
time2 = 0.0061664581298828125
sum of weights now :0.999999999999999
############################################################
Iteration 1 out of 20
processing 4 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 4 K-points on 16 processes: 4.1586 ; per K-point 1.0396 ; proc-sec per K-point 16.6343
factors changed for old points : {2: np.float64(-0.0008)}
time1 = 0.0002770423889160156
time2 = 0.007733583450317383
sum of weights now :0.9999999999999989
############################################################
Iteration 2 out of 20
processing 4 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 4 K-points on 16 processes: 4.4218 ; per K-point 1.1054 ; proc-sec per K-point 17.6872
factors changed for old points : {3: np.float64(-0.0008)}
time1 = 0.0005359649658203125
time2 = 0.007903814315795898
sum of weights now :0.9999999999999988
############################################################
Iteration 3 out of 20
processing 4 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 4 K-points on 16 processes: 4.1847 ; per K-point 1.0462 ; proc-sec per K-point 16.7388
factors changed for old points : {5: np.float64(-0.0008)}
time1 = 0.0002715587615966797
time2 = 0.0069580078125
sum of weights now :0.9999999999999987
############################################################
Iteration 4 out of 20
processing 8 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 8 K-points on 16 processes: 5.7890 ; per K-point 0.7236 ; proc-sec per K-point 11.5780
factors changed for old points : {1: np.float64(-0.0008), 6: np.float64(-0.0008)}
time1 = 0.00024962425231933594
time2 = 0.00768589973449707
sum of weights now :0.9999999999999983
############################################################
Iteration 5 out of 20
processing 12 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 12 K-points on 16 processes: 7.5137 ; per K-point 0.6261 ; proc-sec per K-point 10.0183
factors changed for old points : {9: np.float64(-0.0008), 16: np.float64(-0.0016)}
time1 = 0.0006051063537597656
time2 = 0.008455514907836914
sum of weights now :0.9999999999999983
############################################################
Iteration 6 out of 20
processing 4 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 4 K-points on 16 processes: 4.3441 ; per K-point 1.0860 ; proc-sec per K-point 17.3763
factors changed for old points : {10: np.float64(-0.0008)}
time1 = 0.0002658367156982422
time2 = 0.007470130920410156
sum of weights now :0.9999999999999982
############################################################
Iteration 7 out of 20
processing 4 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 4 K-points on 16 processes: 3.9972 ; per K-point 0.9993 ; proc-sec per K-point 15.9888
factors changed for old points : {8: np.float64(-0.0008)}
time1 = 0.0002560615539550781
time2 = 0.006428241729736328
sum of weights now :0.9999999999999981
############################################################
Iteration 8 out of 20
processing 4 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 4 K-points on 16 processes: 4.2370 ; per K-point 1.0592 ; proc-sec per K-point 16.9479
factors changed for old points : {7: np.float64(-0.0008)}
time1 = 0.0002956390380859375
time2 = 0.006402492523193359
sum of weights now :0.999999999999998
############################################################
Iteration 9 out of 20
processing 4 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 4 K-points on 16 processes: 4.2375 ; per K-point 1.0594 ; proc-sec per K-point 16.9499
factors changed for old points : {4: np.float64(-0.0008)}
time1 = 0.00022912025451660156
time2 = 0.005945682525634766
sum of weights now :0.9999999999999978
############################################################
Iteration 10 out of 20
processing 12 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 12 K-points on 16 processes: 7.0716 ; per K-point 0.5893 ; proc-sec per K-point 9.4289
factors changed for old points : {11: np.float64(-0.0008), 63: np.float64(-0.0016)}
time1 = 0.0003402233123779297
time2 = 0.007120370864868164
sum of weights now :0.9999999999999974
############################################################
Iteration 11 out of 20
processing 8 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 8 K-points on 16 processes: 5.4337 ; per K-point 0.6792 ; proc-sec per K-point 10.8674
factors changed for old points : {12: np.float64(-0.0008), 321: np.float64(-0.0016)}
time1 = 0.0005145072937011719
time2 = 0.006630659103393555
sum of weights now :0.9999999999999972
############################################################
Iteration 12 out of 20
processing 12 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 12 K-points on 16 processes: 7.5277 ; per K-point 0.6273 ; proc-sec per K-point 10.0369
factors changed for old points : {322: np.float64(-0.0016), 466: np.float64(-0.0004)}
time1 = 0.0005013942718505859
time2 = 0.0072078704833984375
sum of weights now :0.9999999999999968
############################################################
Iteration 13 out of 20
processing 12 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 12 K-points on 16 processes: 7.2422 ; per K-point 0.6035 ; proc-sec per K-point 9.6562
factors changed for old points : {37: np.float64(-0.0016), 314: np.float64(-0.0016)}
time1 = 0.00035643577575683594
time2 = 0.006128787994384766
sum of weights now :0.9999999999999967
############################################################
Iteration 14 out of 20
processing 8 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 8 K-points on 16 processes: 5.5168 ; per K-point 0.6896 ; proc-sec per K-point 11.0336
factors changed for old points : {44: np.float64(-0.0016)}
time1 = 0.0003409385681152344
time2 = 0.006665945053100586
sum of weights now :0.9999999999999961
############################################################
Iteration 15 out of 20
processing 12 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 12 K-points on 16 processes: 7.2202 ; per K-point 0.6017 ; proc-sec per K-point 9.6270
factors changed for old points : {88: np.float64(-0.0032000000000000006), 318: np.float64(-0.0016)}
time1 = 0.0005412101745605469
time2 = 0.00928807258605957
sum of weights now :0.9999999999999957
############################################################
Iteration 16 out of 20
processing 12 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 12 K-points on 16 processes: 7.7491 ; per K-point 0.6458 ; proc-sec per K-point 10.3322
factors changed for old points : {69: np.float64(-0.0008), 145: np.float64(-0.0032000000000000006)}
time1 = 0.0003876686096191406
time2 = 0.0058574676513671875
sum of weights now :0.9999999999999951
############################################################
Iteration 17 out of 20
processing 12 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 12 K-points on 16 processes: 7.3091 ; per K-point 0.6091 ; proc-sec per K-point 9.7454
factors changed for old points : {147: np.float64(-0.0032000000000000006), 317: np.float64(-0.0016)}
time1 = 0.0004031658172607422
time2 = 0.007190704345703125
sum of weights now :0.9999999999999946
############################################################
Iteration 18 out of 20
processing 18 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
16 9.0 1.1 10.1
time for processing 18 K-points on 16 processes: 10.2247 ; per K-point 0.5680 ; proc-sec per K-point 9.0886
factors changed for old points : {0: np.float64(-0.0004), 46: np.float64(-0.0016), 219: np.float64(-0.0032000000000000006)}
time1 = 0.0003845691680908203
time2 = 0.0067365169525146484
sum of weights now :0.9999999999999936
############################################################
Iteration 19 out of 20
processing 24 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
16 10.9 5.4 16.3
time for processing 24 K-points on 16 processes: 12.4691 ; per K-point 0.5195 ; proc-sec per K-point 8.3127
factors changed for old points : {90: np.float64(-0.0032000000000000006), 254: np.float64(-0.0032000000000000006), 269: np.float64(-0.0032000000000000006)}
time1 = 0.0006356239318847656
time2 = 0.008974552154541016
sum of weights now :0.9999999999999933
############################################################
Iteration 20 out of 20
processing 8 K points : using 16 processes.
# K-points calculated Wall time (sec) Est. remaining (sec) Est. total (sec)
time for processing 8 K-points on 16 processes: 5.6878 ; per K-point 0.7110 ; proc-sec per K-point 11.3755
factors changed for old points : {307: np.float64(-0.0016)}
time1 = 0.0003428459167480469
Totally processed 589 K-points
run() finished
plot AHC vs energy
[3]:
from matplotlib import pyplot as plt
import numpy as np
theta=0
phi=0
fig, axes = plt.subplots(1, 1, sharey=False, sharex=True, figsize=(10, 8))
axes=[axes]
imin = 0
imax = 20
for i in range(imin, imax+1, 1):
t = (i-imin)/(imax-imin)
res = np.load(f"results/MnTe-soc-y-ahc_int_iter-{i:04d}.npz")
Efermi = res['Energies_0']
ahc = res["data"]
axes[0].plot(Efermi - EF, ahc[:,2]/100, label=f"iter={i}", color=(0, 1-t, t))
axes[0].set_title("AHC internal terms convergence")
# axes[0].set_ylim(-1000, 0)
axes[0].set_xlabel("E-EF (eV)")
axes[0].set_ylabel("AHC (S/cm)")
axes[0].legend()
plt.tight_layout()
plt.savefig("ahc_convergence.png", dpi=300)
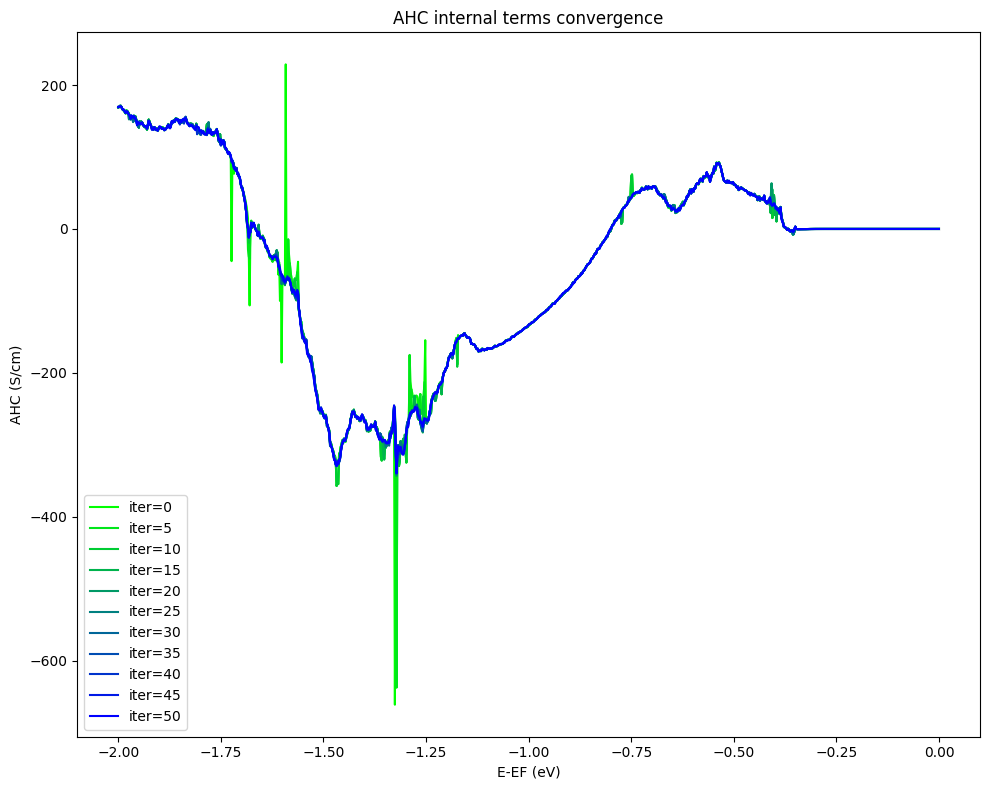
We can observe 3 things, which are generally true, not only for AHC, but for most other effects: 1. The internal terms converge slower than the external ones. 2. The external terms usually give a smaller contribution. Often (but not always) it is negligible. However, check it for a particular material and effect. 3. The external terms take more time to evaluate per k-point.
Thus, it makes sense to evaluate them separately, using different k-point grids and number of iterations for the adaptive integration.
Other directions of magnetic moments
We could try to calculate the AHC when the magnetic moments are along “z” or “x” directions. However, we can see that in those cases the point group contains symmetries which force the AHC to be zero. This can be demonstrated by the following way:
[4]:
from irrep.spacegroup import SpaceGroup
from wannierberri.symmetry.point_symmetry import PointGroup
# x direction
mg = SpaceGroup.from_cell(real_lattice=lattice, positions=positions, typat=[1,1,2,2], magmom=[[1,0,0],[-1,0,0],[0,0,0],[0,0,0]])
# mg.show()
pointgroup = PointGroup(spacegroup=mg)
print (f"components of AHC for magnetic moments along x direction: {pointgroup.get_symmetric_components(rank=1, TRodd=True, Iodd=False)}")
mg = SpaceGroup.from_cell(real_lattice=lattice, positions=positions, typat=[1,1,2,2], magmom=[[0,1,0],[0,-1,0],[0,0,0],[0,0,0]])
# mg.show()
pointgroup = PointGroup(spacegroup=mg)
print (f"components of AHC for magnetic moments along y direction: {pointgroup.get_symmetric_components(rank=1, TRodd=True, Iodd=False)}")
mg = SpaceGroup.from_cell(real_lattice=lattice, positions=positions, typat=[1,1,2,2], magmom=[[0,0,1],[0,0,-1],[0,0,0],[0,0,0]])
# mg.show()
pointgroup = PointGroup(spacegroup=mg)
print (f"components of AHC for magnetic moments along z direction: {pointgroup.get_symmetric_components(rank=1, TRodd=True, Iodd=False)}")
components of AHC for magnetic moments along x direction: ['0=x=y=z']
components of AHC for magnetic moments along y direction: ['0=x=y', 'z']
components of AHC for magnetic moments along z direction: ['0=x=y=z']
[ ]: